Research Articles
Decoding Complexity: A Practical Guide to AI-Driven Epigenetic and Non-Coding RNA Analysis
This article provides a comprehensive guide for biomedical researchers on leveraging artificial intelligence (AI) to analyze epigenetic modifications (e.g., DNA methylation, histone marks) and non-coding RNA (ncRNA) data.
From Bisulfite to Nanopores: A 2024 Guide to 5-Methylcytosine Detection Methods for Precision Epigenetics
This article provides a comprehensive overview of established and emerging methods for detecting 5-methylcytosine (5mC), the cornerstone epigenetic DNA modification.
Integrating 16S rRNA Sequencing, Shotgun Metagenomics, and Host Epigenome Analysis: A Comprehensive Guide for Translational Researchers
This article provides a detailed framework for researchers integrating microbiome profiling (16S rRNA sequencing and shotgun metagenomics) with host epigenome analysis.
Epigenetic Age Acceleration: A Comprehensive 14-Clock Analysis of 174 Disease Associations for Precision Medicine
This systematic review and meta-analysis synthesizes the most current evidence from 14 established epigenetic clocks (e.g., Horvath, Hannum, PhenoAge, GrimAge) and their associations with 174 distinct disease outcomes.
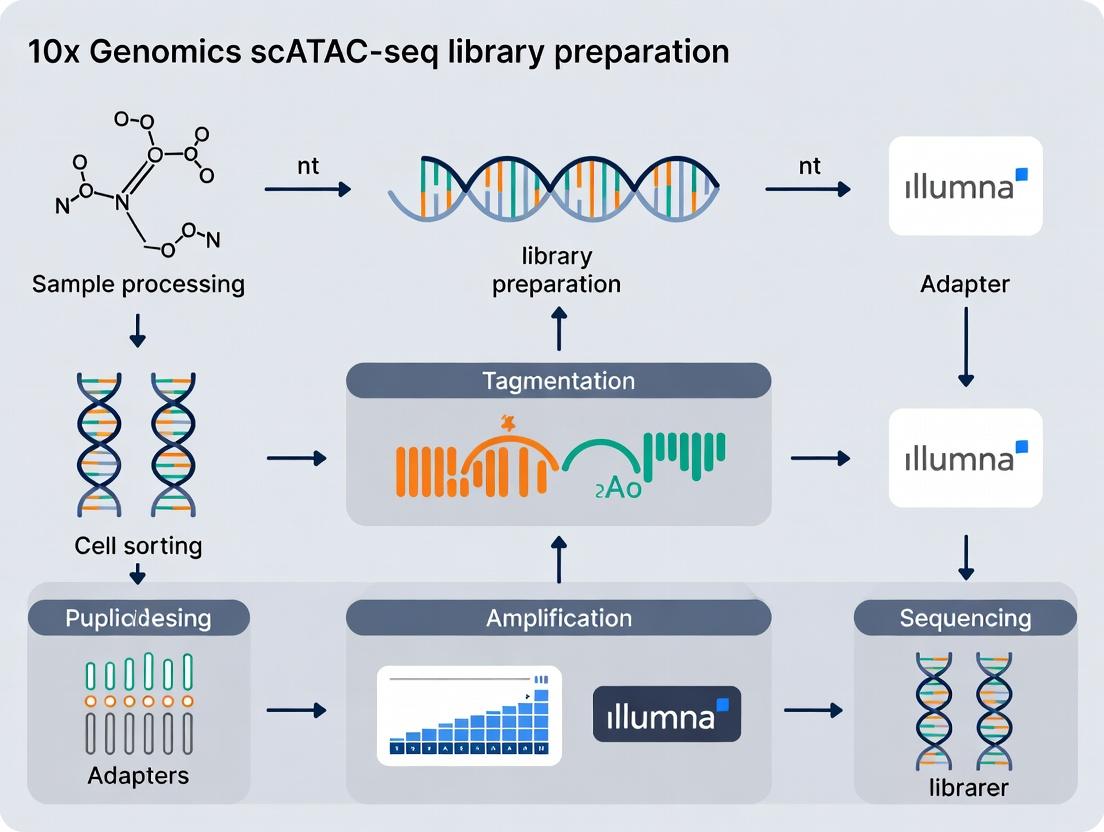
The Complete Guide to 10x Genomics scATAC-seq: From Library Prep to Data Interpretation
This comprehensive guide details the 10x Genomics Chromium Single Cell ATAC solution for profiling chromatin accessibility at single-cell resolution.
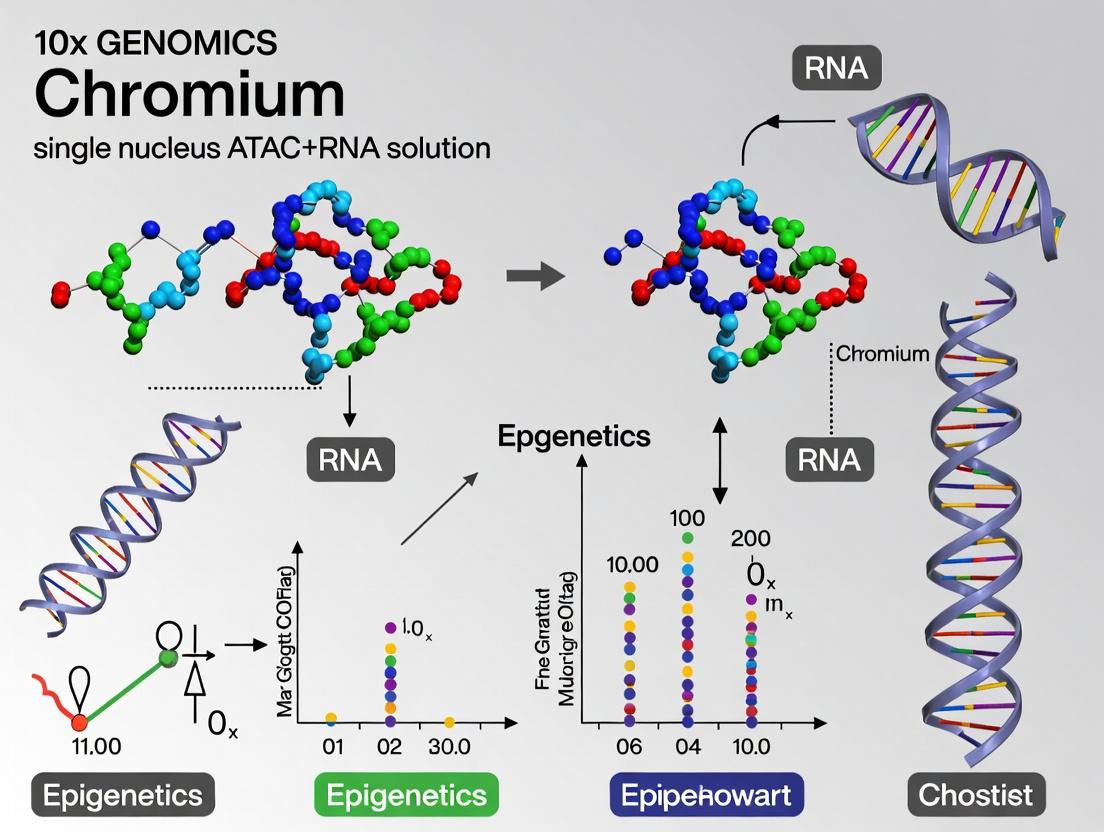
The Ultimate Guide to 10x Genomics Chromium Single Cell Multiome ATAC + Gene Expression: From Protocol to Profiling
This comprehensive guide provides researchers, scientists, and drug development professionals with an in-depth analysis of the 10x Genomics Chromium Single Cell Multiome ATAC + Gene Expression platform.
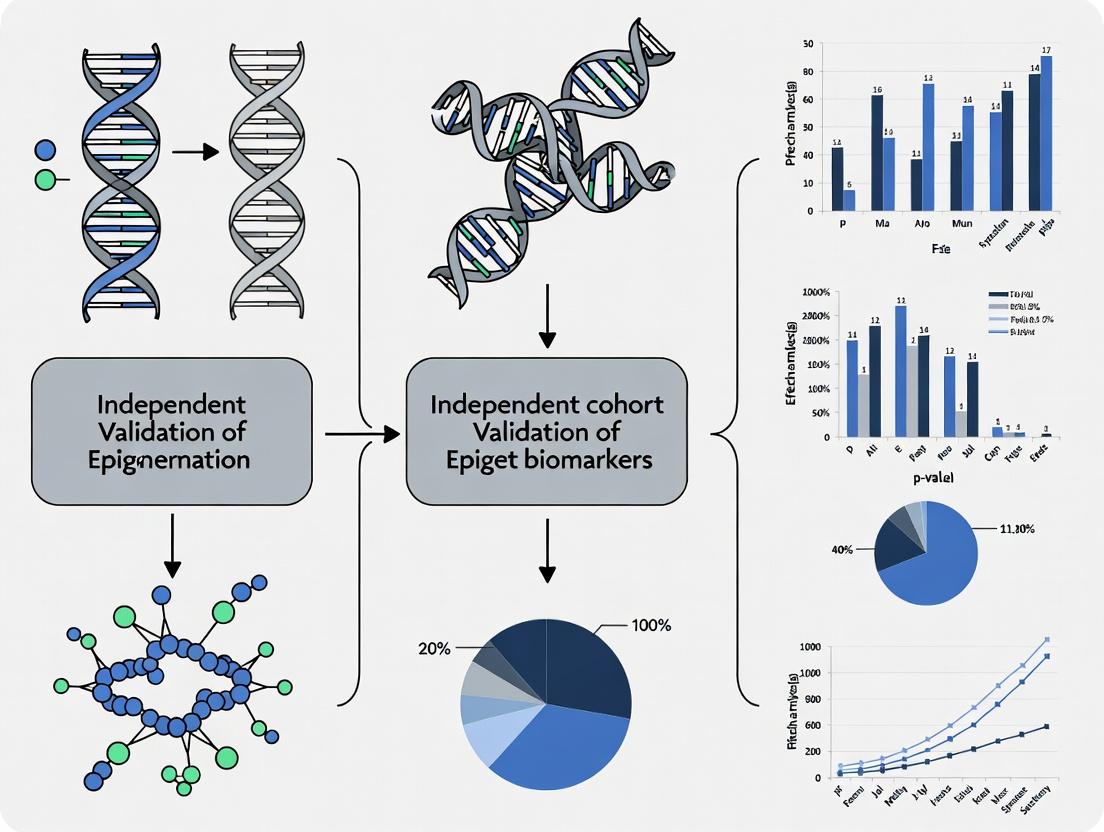
Beyond Discovery: A Practical Guide to Independent Cohort Validation of Epigenetic Biomarkers
For researchers and drug development professionals, translating promising epigenetic biomarker discoveries into robust, clinically useful tools requires rigorous independent validation.
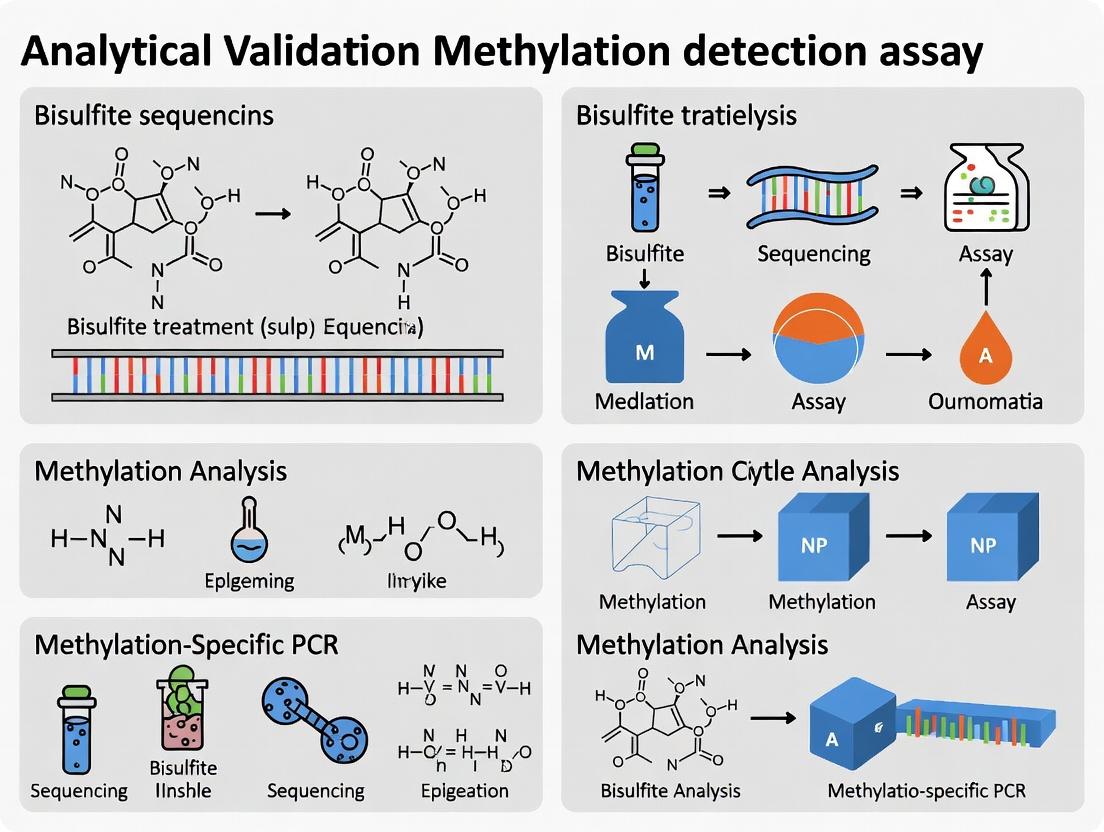
The Definitive Guide to Analytical Validation for DNA Methylation Detection Assays: From Design to Implementation
This comprehensive guide details the critical framework for the analytical validation of DNA methylation detection assays, essential for biomarker discovery and clinical diagnostics.
Beyond a Single Disease: The Critical Role of Cross-Cancer Validation in Epigenetic Biomarker Discovery
This article examines the necessity and methodologies for the cross-cancer validation of epigenetic signatures, focusing on DNA methylation patterns.
Beyond Promise: Evaluating Epigenetic Biomarkers in Prospective Clinical Trials for Precision Medicine
Epigenetic biomarkers, particularly DNA methylation signatures and cell-free nucleosome profiles, offer immense potential for early disease detection, prognosis, and monitoring therapeutic response.