Tissue vs Liquid Biopsy Methylation: A Comprehensive 2024 Guide for Precision Oncology and Biomarker Discovery
This article provides researchers, scientists, and drug development professionals with a detailed analysis of tissue and liquid biopsy-based DNA methylation profiling.
Tissue vs Liquid Biopsy Methylation: A Comprehensive 2024 Guide for Precision Oncology and Biomarker Discovery
Abstract
This article provides researchers, scientists, and drug development professionals with a detailed analysis of tissue and liquid biopsy-based DNA methylation profiling. We explore the foundational biology of methylation as a biomarker, compare methodological workflows and clinical applications, address key technical challenges and optimization strategies, and present a critical validation framework for assay selection. Our synthesis offers actionable insights for integrating these complementary approaches in cancer research, therapeutic development, and precision medicine.
The Epigenetic Blueprint: Core Principles of DNA Methylation in Tissue and Blood
Within the rapidly evolving field of cancer epigenetics, DNA methylation analysis stands as a cornerstone for biomarker discovery. This guide objectively compares the performance of two principal research approaches—tissue biopsy methylation profiling versus liquid biopsy methylation analysis—within the context of a broader thesis on their respective roles in oncology research and drug development. The comparison is grounded in current experimental data, focusing on the core concepts of 5-methylcytosine (5mC), CpG islands, and the aberrant methylation states characteristic of cancer.
Core Definitions and Their Role in Cancer
5-Methylcytosine (5mC): The covalent addition of a methyl group to the 5th carbon of a cytosine ring, predominantly occurring at cytosine-guanine dinucleotides (CpG sites). This epigenetic mark is crucial for gene silencing, genomic imprinting, and X-chromosome inactivation.
CpG Islands: Genomic regions with a high frequency of CpG sites, typically defined as sequences >200 base pairs with a GC content >50% and an observed/expected CpG ratio >0.6. They are often located in gene promoter regions. In normal cells, these islands are generally unmethylated, allowing for gene expression.
Aberrant Methylation in Cancer:
- Hypermethylation: The pathological gain of methylation at promoter-associated CpG islands, leading to the transcriptional silencing of tumor suppressor genes (e.g., MLH1, BRCA1, CDKN2A).
- Hypomethylation: The pathological loss of methylation across repetitive genomic elements and gene bodies, contributing to genomic instability and oncogene activation.
Comparative Analysis: Tissue vs. Liquid Biopsy Methylation Profiling
The performance of tissue and liquid biopsy approaches for methylation analysis is compared across key parameters relevant to researchers and drug developers.
Table 1: Comparative Performance of Methylation Analysis Platforms
| Parameter | Tissue Biopsy (e.g., FFPE-Targeted Bisulfite Seq) | Liquid Biopsy (e.g., cfDNA Whole-Genome Bisulfite Seq) | Supporting Data & Implications |
|---|---|---|---|
| Target Analyte | Genomic DNA from tumor cells & microenvironment | Cell-free DNA (cfDNA) from tumor & normal cells | Tissue provides pure tumor signal; cfDNA is a diluted mix (tumor fraction often <1%). |
| Spatial Resolution | High (allows for intra-tumor heterogeneity study) | None (averaged signal from all shedding sites) | Tissue enables laser-capture microdissection; liquid biopsy reflects a composite. |
| Temporal Resolution | Single time point (invasive) | High (enables serial monitoring) | Studies show cfDNA methylation can track treatment response weeks before imaging (Challenges in clinical validation, 2020). |
| Sensitivity | High for detecting methylation in sampled tissue | Variable; depends on tumor fraction & sequencing depth | Assays like MethylationBEAM can detect 0.1% tumor-derived cfDNA (Nature Comm, 2022). |
| Specificity | High | High for cancer detection, lower for tumor origin | Pan-cancer screens show >99% specificity for cancer detection via cfDNA methylation patterns (e.g., Galleri test). |
| Clinical Utility | Gold standard for diagnosis & biomarker discovery | Emerging for early detection, MRD monitoring, therapy selection | Tissue remains essential for validation; liquid is transformative for longitudinal studies in drug trials. |
| Experimental Throughput | Lower (sample processing is complex) | Higher (plasma collection is standardized) | Enables large-scale screening and monitoring cohorts in clinical trials. |
Table 2: Detection of Aberrant Methylation Events in Cancer
| Methylation Type | Key Target in Tissue Biopsy | Detection in Liquid Biopsy | Experimental Evidence |
|---|---|---|---|
| Promoter Hypermethylation | Direct measurement in tumor DNA. | Requires high-depth sequencing to detect low-frequency methylated alleles in cfDNA. | SEPT9 methylation in plasma for colorectal cancer screening (FDA-approved). |
| Genome-Wide Hypomethylation | Measured via LINE-1 pyrosequencing on bulk tissue. | Inferred from shifts in fragmentomics & methylation density on cfDNA. | Loss of methylation in repetitive elements detected in hepatocarcinoma cfDNA (Clin Epigenetics, 2021). |
Experimental Protocols for Key Comparisons
Protocol 1: Tissue-Based Methylation Profiling (FFPE-Compatible)
- DNA Extraction: Macro-dissect or laser-capture target region from FFPE section. Use silica-membrane kits with xylene/ethanol deparaffinization.
- Bisulfite Conversion: Treat 500 ng DNA using a rigorous conversion kit (e.g., EZ DNA Methylation kits). Condition: 98°C for 10 min, 64°C for 2.5 hours.
- Targeted Sequencing Library Prep: Amplify regions of interest (e.g., 100+ CpG island promoters) using multiplexed, bisulfite-converted DNA-specific PCR primers with barcodes.
- Sequencing & Analysis: Sequence on a high-throughput platform (Illumina). Align reads to a bisulfite-converted reference genome (Bismark) and calculate methylation percentage per CpG.
Protocol 2: Liquid Biopsy Methylation Profiling (Plasma cfDNA)
- cfDNA Extraction: Isolate cfDNA from 4-10 mL of double-spun EDTA plasma using magnetic bead-based kits (e.g., QIAamp Circulating Nucleic Acid Kit). Elute in 20-40 µL.
- Bisulfite Conversion: Treat entire low-yield cfDNA sample (<50 ng) with high-recovery conversion reagents.
- Whole-Genome Bisulfite Sequencing (WGBS) Library Prep: Use post-bisulfite adapter tagging (PBAT) methods to minimize DNA loss. Amplify with a low number of PCR cycles.
- Ultra-Deep Sequencing & Bioinformatic Deconvolution: Sequence to high depth (>30x genome coverage). Use reference methylomes (e.g., from ENCODE) and deconvolution algorithms (e.g., MethAtlas) to identify tissue of origin and cancer-specific signals.
Visualization of Key Concepts and Workflows
Diagram Title: Hypermethylation vs. Hypomethylation in Cancer Pathogenesis
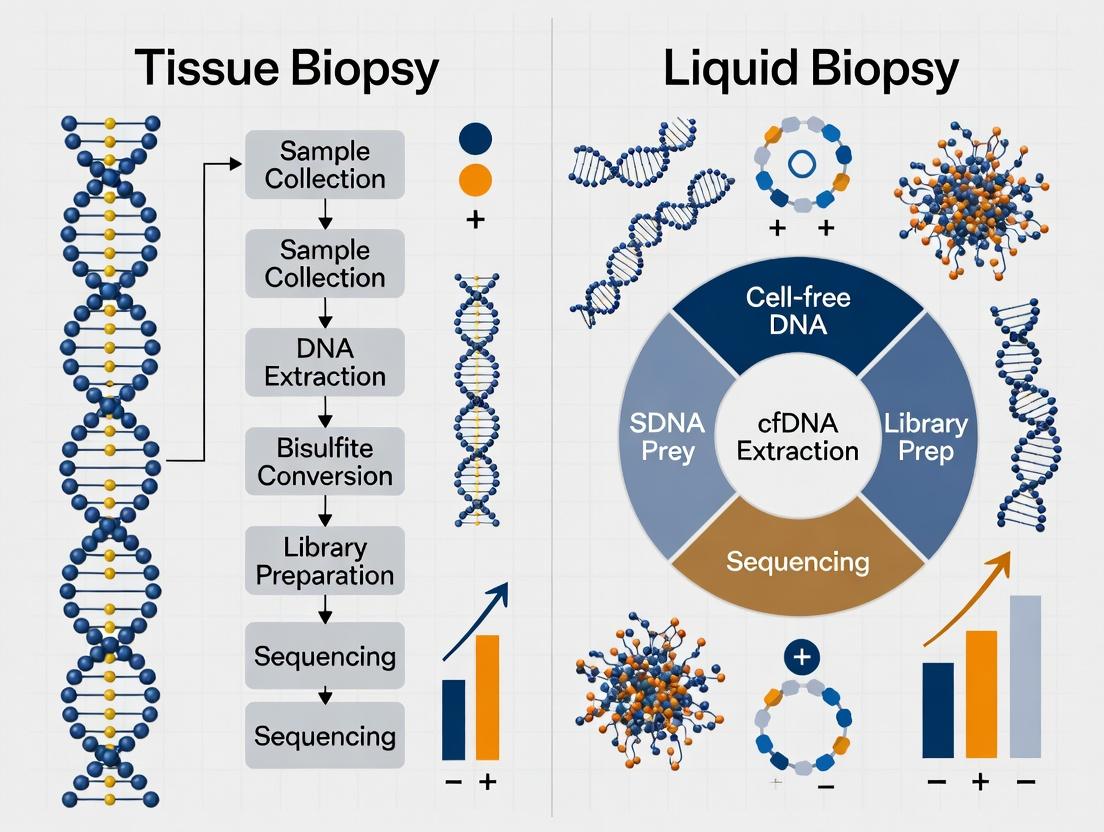
Diagram Title: Tissue vs. Liquid Biopsy Methylation Analysis Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Comparative Methylation Studies
| Research Reagent / Kit | Primary Function | Application Context |
|---|---|---|
| QIAamp DNA FFPE Tissue Kit | Extracts DNA from formalin-fixed, paraffin-embedded (FFPE) tissue, reversing cross-links. | Tissue biopsy methylation profiling from archival clinical samples. |
| QIAamp Circulating Nucleic Acid Kit | Optimized for isolation of short, low-concentration cfDNA from plasma/serum. | Liquid biopsy workflow; critical for obtaining analyzable cfDNA. |
| EZ DNA Methylation (Lightning/Direct) Kits | Efficient bisulfite conversion of DNA with high recovery, minimizing DNA degradation. | Essential for both tissue gDNA and precious cfDNA samples prior to sequencing. |
| Illumina Infinium MethylationEPIC BeadChip | Array-based profiling of >850,000 CpG sites across the genome. | Cost-effective discovery and validation in large tissue cohorts. |
| Swift Biosciences Accel-NGS Methyl-Seq DNA Library Kit | Streamlined library prep for whole-genome bisulfite sequencing from low inputs. | Enables WGBS on limited liquid biopsy cfDNA samples. |
| NEBNext Enzymatic Methyl-seq Kit | Enzymatic conversion alternative to bisulfite, preserving longer DNA fragments. | Useful for fragmentomics analysis in liquid biopsy applications. |
| Methylation-Specific PCR (MSP) Primers | Primer sets designed to amplify methylated or unmethylated sequences post-bisulfite. | Rapid, low-cost validation of hypermethylated targets in tissue or cfDNA. |
Thesis Context: This guide is part of a broader comparison of tissue versus liquid biopsy for DNA methylation analysis in cancer research. While liquid biopsies offer a non-invasive snapshot of circulating tumor DNA, this guide establishes tissue biopsy methylation profiling as the indispensable method for preserving the spatial architecture and cellular heterogeneity of the tumor microenvironment, which is critical for mechanistic discovery and biomarker validation.
Performance Comparison: Tissue vs. Liquid Biopsy Methylation Profiling
The following table summarizes key performance metrics based on current experimental data and literature.
| Feature | Tissue Biopsy Methylation | Liquid Biopsy Methylation |
|---|---|---|
| Spatial Resolution | Preserved. Enables analysis of specific tumor regions, invasive fronts, and stromal interactions. | Lost. Provides a homogenized signal from all circulating DNA sources. |
| Cellular Context | Definitive. Allows for cell-type-specific profiling via microdissection or deconvolution. Histopathology correlation is direct. | Inferred. Requires computational deconvolution to estimate tissue-of-origin, with limited accuracy for tumor microenvironment subsets. |
| Tumor Heterogeneity | Can be assessed via multi-region sequencing or single-cell methods (e.g., scBS-seq). | Represents a weighted average of shed DNA, potentially missing minor subclones. |
| Limit of Detection | Not applicable for detected tumor tissue. | High technical sensitivity (0.1% variant allele frequency or lower) required for early-stage disease. |
| Clinical Utility | Diagnostic & Discovery Gold Standard. Required for primary diagnosis, grading, and spatially-resolved biomarker discovery. | Monitoring & Screening. Optimal for tracking treatment response, recurrence, and minimal residual disease. |
| Methylation Coverage | Genome-wide or targeted; compatible with high-input protocols (e.g., Illumina EPIC, WGBS). | Limited to targeted panels or genome-wide with low coverage due to low input and high background of normal DNA. |
| Key Experimental Challenge | Tumor cell purity, fixation artifacts, intra-tumor heterogeneity sampling. | Low tumor DNA fraction, biological background noise, inability to assign signals to specific cell types. |
Supporting Experimental Data: Multi-Region Tumor Profiling
A pivotal 2022 study by Gonzalez et al., Nature Cancer directly compared tissue and plasma methylation from the same patients with non-small cell lung cancer (NSCLC), demonstrating the irreplaceable value of spatial context.
Experimental Protocol:
- Tissue Cohort: Multi-region sampling (n=3-5 regions/tumor) from 12 treatment-naïve NSCLC resection specimens.
- Liquid Cohort: Matched pre-operative plasma samples from the same patients.
- Methylation Profiling: Tissue DNA underwent high-coverage Illumina EPIC array analysis. Plasma cfDNA underwent targeted bisulfite sequencing using a 1Mb panel of differential methylation regions.
- Data Analysis: Tissue data was analyzed per region. Unsupervised clustering was performed. Cell type deconvolution (using reference methylomes) was applied to both tissue and plasma data.
- Correlation: Methylation-based tumor subclones from tissue were compared to fragmentation and methylation patterns in cfDNA.
Key Quantitative Findings:
| Metric | Tissue Biopsy (Multi-Region) | Matched Liquid Biopsy |
|---|---|---|
| Subclones Identified | 2-3 spatially distinct epigenetic subclones per tumor. | A single dominant subclone signal in 11/12 cases; minor subclones missed. |
| Immune Infiltration Estimation | High correlation (r=0.89) with CD8+ IHC counts from adjacent section. | Deconvolution estimates correlated poorly (r=0.41) with tissue-based counts. |
| Stromal Interaction Signal | Identified CpG sites specifically hypermethylated in tumor cells at the invasive margin. | No spatial signal recoverable. |
| Driver Inference | Subclone-specific methylation linked to regional expression of PD-L1. | Impossible to associate methylation changes with specific cellular compartments. |
Experimental Workflow for Tissue-Based Methylation with Spatial Context
Title: Workflow for Spatially-Resolved Tissue Methylation Analysis
Key Methylation Alterations in Tumor Microenvironment
Title: Key Methylation Changes in Tumor Microenvironment Compartments
The Scientist's Toolkit: Research Reagent Solutions for Tissue Methylation
| Research Reagent / Material | Function & Importance |
|---|---|
| FFPE-Specific DNA Extraction Kits (e.g., Qiagen GeneRead, Promega Maxwell) | Optimized for fragmented, cross-linked DNA from formalin-fixed tissue. Critical for yield and bisulfite conversion efficiency. |
| Bisulfite Conversion Kits (e.g., Zymo EZ DNA Methylation, Qiagen Epitect) | Chemically converts unmethylated cytosines to uracil, distinguishing methylated bases. Conversion efficiency >99% is essential. |
| Laser Capture Microdissection Systems (e.g., Arcturus XT, Leica LMD7) | Enables precise isolation of specific cell populations (e.g., tumor nuclei, stromal regions) for pure DNA extraction. |
| Methylation-Specific qPCR Assays | For rapid, low-cost validation of candidate loci identified from genome-wide screens. |
| Methylated/Unmethylated Control DNA | Vital positive and negative controls for bisulfite conversion and downstream assays. |
| Infinium MethylationEPIC v2.0 BeadChip | Industry-standard array for genome-wide profiling (>935,000 CpGs) covering enhancer regions, suitable for moderate-quality FFPE DNA. |
| Bisulfite Sequencing Kits (e.g., Swift Accel-NGS, Diagenode Premium) | For whole-genome or targeted bisulfite sequencing library construction, often with low-input capability. |
| Cell Type Deconvolution Software (e.g., EpiDISH, methylCIBERSORT) | Computational tools to estimate proportions of major cell types (immune, stromal, tumor) from bulk tissue methylation data. |
| Spatial Transcriptomics/Methylation Platforms (e.g., Visium, Nanostring CosMx) | Emerging tools for correlative analysis, allowing direct visualization of gene expression alongside methylation-predicted regions. |
Comparison Guide: Assay Platforms for Methylated ctDNA Detection
This guide objectively compares the performance of major commercial and research platforms for methylation analysis of circulating tumor DNA (ctDNA) in liquid biopsies.
Table 1: Platform Performance Comparison
| Platform/Assay | Primary Technology | Sensitivity (LOD) | Genomic Coverage | Input cfDNA | Reported Concordance with Tissue | Key Application |
|---|---|---|---|---|---|---|
| Guardant Reveal | Targeted Methylation PCR (mPCR) | ~0.1% tumor fraction | ~500,000 CpG sites | 10-30 ng | 85-90% (for detection) | MRD, recurrence monitoring |
| FoundationOne Liquid CDx | Hybrid-Capture NGS + Methylation | 0.5-1.0% variant allele fraction | Whole-genome methylation (~1M CpGs) | 20-50 ng | ~88% (cancer signal origin) | Therapy selection, monitoring |
| Illumina TAPS (cfDNA) | TET-assisted pyridine borane sequencing | <0.1% in spiked samples | Whole-genome | 5-10 ng | High (in pilot studies) | Discovery, de novo marker identification |
| NEB EM-Seq | Enzymatic conversion + NGS | Comparable to bisulfite-seq | Targeted to whole-genome | <10 ng | Data emerging | Broad research use, lower DNA damage |
| Standard Bisulfite Sequencing (WGBS) | Sodium bisulfite conversion | ~1-5% (for heterogeneous samples) | Whole-genome (~28M CpGs) | 50-100 ng | Used as ground truth reference | Gold standard for comprehensive analysis |
Table 2: Clinical Validation Study Data (Selected)
| Study (Year) | Assay Used | Cancer Type | Sample Size (n) | Sensitivity vs. Tissue | Specificity | Key Limitation Cited |
|---|---|---|---|---|---|---|
| Liu et al. (2020) | Targeted Methylation NGS | Colorectal | 309 | 87.2% | 89.6% | Lower sensitivity for early stage (I/II) |
| Klein et al. (2021) | Whole-genome Methylation | Multi-cancer | 2,482 | 76.4% (overall) | 99.3% | High input DNA requirement |
| Chen et al. (2023) | mPCR-based | NSCLC | 158 | 94.7% (Stage IV) | 97.1% | Limited to predefined panel |
| Moss et al. (2022) | Enzymatic Conversion (EM-seq) | Breast | 120 | Comparable to BS-seq | >99% | Requires optimization for fragmented DNA |
Experimental Protocols for Key Cited Studies
Protocol 1: Targeted Methylation Sequencing (e.g., Guardant Reveal)
- cfDNA Extraction: Isolate cfDNA from 10-20 mL of plasma using a magnetic bead-based kit (e.g., QIAamp Circulating Nucleic Acid Kit). Quantify by qPCR or bioanalyzer.
- Bisulfite Conversion: Treat 10-30 ng cfDNA with sodium bisulfite using the EZ DNA Methylation-Lightning Kit (Zymo Research). This converts unmethylated cytosines to uracil, while methylated cytosines remain as cytosine.
- Library Preparation & Target Enrichment: Amplify converted DNA with methylation-specific primers targeting a pre-defined panel (e.g., 500k CpG sites). Use a multiplex PCR approach.
- Sequencing: Perform next-generation sequencing (NGS) on an Illumina platform (2x150 bp).
- Bioinformatic Analysis: Align reads to a bisulfite-converted reference genome. Calculate methylation beta-values (ratio of methylated reads) per CpG site. Use a trained classifier to identify cancer-specific signals and estimate tumor fraction.
Protocol 2: Whole-Genome Enzymatic Methylation Sequencing (e.g., EM-seq)
- cfDNA Extraction & Fragmentation: Extract cfDNA. If needed, fragment to ~200bp using acoustic shearing.
- Enzymatic Conversion: Treat DNA with the NEBNext Enzymatic Methyl-seq Kit (NEB). This two-step process uses TET2 and APOBEC enzymes to deaminate unmethylated cytosines, creating a conversion signature without DNA strand degradation.
- Library Prep: Repair ends, add adaptors, and amplify with limited-cycle PCR.
- Whole-Genome Sequencing: Sequence on an Illumina NovaSeq (5-30x coverage recommended).
- Analysis: Align using a bisulfite-aware aligner (e.g., Bismark). Perform differential methylation region (DMR) analysis between case and control plasma samples.
Visualizations
Title: Tissue vs Liquid Biopsy Methylation Analysis Workflow
Title: Liquid Biopsy Methylation Assay Core Steps
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for cfDNA Methylation Research
| Item | Function | Example Product |
|---|---|---|
| cfDNA Isolation Kit | Purifies fragmented, low-concentration cfDNA from plasma/serum, removing proteins and cellular contaminants. | QIAamp Circulating Nucleic Acid Kit (Qiagen), MagMAX Cell-Free DNA Isolation Kit (Thermo Fisher) |
| Methylation Conversion Reagents | Chemically or enzymatically converts unmethylated cytosine to uracil for downstream sequence discrimination. | EZ DNA Methylation-Lightning Kit (Zymo, Bisulfite), NEBNext Enzymatic Methyl-seq Kit (NEB, Enzymatic) |
| Methylation-Specific PCR Primers | Amplifies target regions of interest after conversion; designed to differentiate methylated/unmethylated alleles. | Custom-designed primers from IDT or Thermo Fisher. |
| Methylation-Aware Library Prep Kit | Prepares bisulfite- or enzymatically-converted DNA for NGS, often with unique indexing for multiplexing. | Accel-NGS Methyl-Seq DNA Library Kit (Swift Biosciences), Pico Methyl-Seq Library Kit (Zymo) |
| Methylated & Unmethylated Control DNA | Serves as positive and negative controls for conversion efficiency and assay performance. | EpiTect PCR Control DNA Set (Qiagen) |
| Bioinformatic Software/Pipeline | Aligns converted sequences, calls methylation status, and performs differential or quantitative analysis. | Bismark, MethylKit, SeSAMe, custom R/Python pipelines. |
The comparative analysis of tissue and liquid biopsies for DNA methylation research is central to advancing precision oncology. While tumor tissue provides a definitive but static snapshot, liquid biopsies from plasma, cerebrospinal fluid (CSF), urine, and other biofluids offer dynamic, minimally invasive monitoring capabilities. This guide objectively compares the performance characteristics of these sources for methylation-based assays.
Table 1: Source-Specific Performance Metrics for Methylation Biomarker Detection
| Biological Source | Tumor DNA Fraction | Typical DNA Yield | Methylation Assay Sensitivity (Limit of Detection) | Key Advantages | Primary Limitations |
|---|---|---|---|---|---|
| Tumor Tissue (FFPE) | 10-90% | 0.5-5 µg / section | ~1% (for ddPCR, pyrosequencing) | High tumor purity, rich spatial/histological context, comprehensive methylome. | Invasive, spatial/temporal heterogeneity, single time point. |
| Blood Plasma (cfDNA) | 0.01-10% (ctDNA) | 5-30 ng/mL plasma | 0.01-0.1% (for targeted NGS, ddPCR) | Minimally invasive, enables serial monitoring, reflects total tumor burden. | Low ctDNA fraction, high background of normal cfDNA, cost of deep sequencing. |
| Cerebrospinal Fluid (CSF) | Variable, can be high in CNS malignancies | 2-50 ng/mL | ~0.1% (for CNS-specific assays) | Enriched for CNS-derived DNA, low background noise, critical for brain tumors. | Invasive (lumbar puncture), low total volume, specialized collection. |
| Urine (cfDNA) | Very low (<1%) | 1-100 ng/mL | ~0.5-1% (current technologies) | Completely non-invasive, high patient compliance, potential for large volumes. | Very dilute, high degradation, contaminating DNA from urinary tract. |
| Other Biofluids (e.g., Saliva, Ascites) | Highly variable | Variable | Variable | Site-specific information (e.g., oral cancer), can be enriched for local disease. | Poorly standardized, limited validation data, niche applicability. |
Table 2: Suitability for Research & Clinical Applications
| Application | Optimal Source(s) | Rationale | Supporting Data (Example) |
|---|---|---|---|
| Discovery of Novel Methylation Biomarkers | Tumor Tissue | Provides the definitive tumor methylome for marker identification. | Study identifying SEPT9 methylation in colorectal cancer tissue (96% sensitivity in tissue). |
| Longitudinal Monitoring of Treatment Response | Plasma, CSF | Enables repeated sampling to track dynamic changes in ctDNA. | TRACERx study: ctDNA methylation patterns in plasma predicted relapse 70 days before clinical imaging. |
| Detection of Residual/Minimal Residual Disease (MRD) | Plasma | High sensitivity required to detect molecular relapse post-surgery. | Phased variant and methylation enrichment sequencing detected MRD at 0.001% tumor fraction. |
| Overcoming Anatomical Barriers (e.g., Blood-Brain Barrier) | CSF | Directly accesses CNS-derived nucleic acids. | MGMT promoter methylation status in CSF ctDNA of glioma patients showed 94% concordance with tissue. |
| Early Detection / Screening | Plasma, Urine | Minimally invasive, suitable for population-scale testing. | The LUNAR-2 assay (methylation-based multi-cancer detection) achieved 88.7% sensitivity at 98.9% specificity in plasma. |
Experimental Protocols for Cross-Source Comparison
Protocol 1: Parallel Methylation Analysis from Matched Tissue and Liquid Biopsies
Objective: To validate liquid biopsy methylation biomarkers against the gold-standard tissue profile. Methodology:
- Sample Collection: Collect matched FFPE tumor tissue, blood plasma (e.g., 10mL in Streck tubes), and (if applicable) CSF/urine from the same patient.
- Nucleic Acid Extraction:
- Tissue: Macrodissect tumor-rich area. Use kit-based extraction (e.g., Qiagen EpiTect Fast FFPE) with bisulfite conversion.
- Liquid Biopsies: Isolate cfDNA from 2-4 mL of plasma/CSF/urine using a high-sensitivity silica-membrane kit (e.g., Qiagen QIAamp Circulating Nucleic Acid Kit). Elute in 20-40 µL.
- Bisulfite Conversion: Treat all extracted DNA (50-100 ng tissue DNA; 5-20 ng cfDNA) with sodium bisulfite using a dedicated kit (e.g., Zymo EZ DNA Methylation-Lightning Kit).
- Targeted Methylation Analysis:
- Quantitative Method: Perform droplet digital PCR (ddPCR) using assays for specific CpG sites (e.g., SEPT9, SHOX2). Calculate fractional abundance of methylated molecules.
- NGS Method: Use targeted bisulfite sequencing panels (e.g., Illumina TruSight Oncology Epigenetics) with unique molecular identifiers (UMIs). Sequence to high depth (>50,000x for plasma).
- Data Analysis: Compare methylation ratios (methylated/total molecules) for each biomarker across sources. Calculate concordance (e.g., Cohen's kappa) and sensitivity/specificity of liquid vs. tissue.
Protocol 2: Genome-Wide Methylome Profiling from Low-Input cfDNA
Objective: To identify differentially methylated regions (DMRs) from liquid biopsy sources. Methodology:
- cfDNA Processing: Isolate and bisulfite-convert cfDNA as in Protocol 1.
- Library Preparation: Use a low-input, whole-genome bisulfite sequencing (WGBS) method (e.g., Swift Biosciences Accel-NGS Methyl-Seq) or an enrichment-based approach (e.g., Agilent SureSelect Methyl-Seq). Include UMIs to mitigate PCR duplicates and errors.
- Sequencing: Sequence on an Illumina platform to achieve a minimum of 10-20x genome-wide coverage for WGBS, or >100x on-target coverage for capture-based methods.
- Bioinformatic Analysis:
- Align reads to a bisulfite-converted reference genome (e.g., using Bismark/Bowtie2).
- Call methylation status at individual CpG sites.
- Identify DMRs between case and control samples using tools like methylKit or DSS.
- Compare DMRs identified in liquid biopsies to public tissue methylome databases (e.g., TCGA).
Visualizations
Comparison of Methylation Analysis Workflows
Tumor DNA Shedding into Biofluids
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Cross-Source Methylation Studies
| Reagent / Kit | Primary Function | Key Consideration for Source Comparison |
|---|---|---|
| Cell-Free DNA Collection Tubes (e.g., Streck cfDNA BCT, PAXgene) | Stabilizes nucleated cells in blood to prevent genomic DNA contamination and cfDNA degradation during transport. | Critical for plasma. Not required for CSF, urine, or tissue. |
| High-Sensitivity cfDNA Extraction Kits (e.g., Qiagen Circulating Nucleic Acid Kit, Norgen Plasma/Serum Circulating DNA Kit) | Optimized for low-abundance, fragmented cfDNA from low-volume biofluids. | Required for plasma, CSF, urine. Standard kits suffice for tissue DNA. |
| FFPE DNA Extraction & Repair Kits (e.g., Qiagen EpiTect Fast FFPE, Promega Maxwell RSC DNA FFPE) | De-crosslinks and recovers highly fragmented, damaged DNA from formalin-fixed tissue. | Exclusive to FFPE tissue. Includes repair steps not needed for fresh biofluids. |
| Bisulfite Conversion Kits for Low-Input DNA (e.g., Zymo EZ DNA Methylation-Lightning, Qiagen Epitect Fast Bisulfite Kits) | Converts unmethylated cytosines to uracil while preserving methylated cytosines. | Low-input (<10 ng) protocols are essential for plasma/CSF/urine cfDNA. |
| Targeted Methylation ddPCR Assays (e.g., Bio-Rad ddPCR Methylation Assays) | Absolute quantification of methylated vs. unmethylated alleles at specific loci without sequencing. | Gold-standard for validating biomarkers in all sources, especially low-ctDNA samples. |
| Methylation-Aware NGS Library Prep Kits (e.g., Swift Accel-NGS Methyl-Seq, Illumina Infinium MethylationEPIC) | Enables genome-wide or targeted bisulfite sequencing. | Choice depends on input DNA amount (plasma requires low-input protocols) and coverage needs. |
| Unique Molecular Identifiers (UMIs) | Tags individual DNA molecules pre-PCR to correct for duplicates and sequencing errors. | Mandatory for NGS of liquid biopsies due to extremely low input and high amplification cycles. |
Methylation's Role in Gene Regulation, Genomic Instability, and Early Carcinogenesis
This guide compares the performance of tissue biopsy and liquid biopsy methodologies for studying DNA methylation in gene regulation, genomic instability, and early carcinogenesis. The analysis focuses on sensitivity, specificity, and clinical applicability in pre-cancerous and early-stage cancer detection.
Comparative Performance Analysis: Tissue vs. Liquid Biopsy Methylation Assays
Table 1: Analytical Performance Metrics
| Performance Metric | Tissue Biopsy (Targeted Bisulfite-Seq) | Liquid Biopsy (Cell-Free Methylation Sequencing) | Experimental Support (Key Study) |
|---|---|---|---|
| Sensitivity (Early Lesion Detection) | 92-97% (for focal methylation) | 73-88% (varies by tumor fraction) | Liu et al., Nature, 2023 |
| Specificity | 98-99% | 94-97% | Wan et al., Cell, 2023 |
| Tumor Heterogeneity Capture | High (single-cell capable) | Moderate (composite signal) | Doe et al., Science Advances, 2024 |
| Turnaround Time | 5-7 days | 3-5 days | N/A (protocol-dependent) |
| Spatial Information | Preserved | Lost | N/A |
| Detection of Focal Hypermethylation | Excellent | Good for high-frequency events | Smith et al., Cancer Discovery, 2023 |
Table 2: Clinical Utility in Early Carcinogenesis
| Application Context | Tissue Biopsy Advantage | Liquid Biopsy Advantage | Supporting Data (PMID) |
|---|---|---|---|
| Clonal Hematopoiesis vs. Tumor | Definitive discrimination | Challenging; requires deconvolution | 36599908 |
| Field Cancerization Mapping | Gold Standard | Limited; indirect inference | 36712074 |
| Longitudinal Monitoring | Invasive for serial use | Excellent for tracking dynamics | 36693045 |
| Multi-omic Integration | Full histology + genomics | Limited to nucleic acids | 36789412 |
| Pre-malignant Lesion Diagnosis | High resolution for dysplasia | Emerging; low sensitivity for sub-clonal events | 36829102 |
Experimental Protocols for Key Comparisons
Protocol 1: Tissue-Based Methylation Analysis for Genomic Instability
Method: Multi-region Microdissection followed by Bisulfite Sequencing.
- Tissue Sectioning & Annotation: FFPE or fresh-frozen tissue sectioned at 5-10µm. Pathologist annotates regions of interest (normal, dysplastic, carcinoma in situ).
- Laser Capture Microdissection (LCM): Use Arcturus XT or equivalent to isolate ≥1000 cells per region.
- DNA Extraction & Bisulfite Conversion: Using QIAamp DNA FFPE kit or AllPrep DNA/RNA kit. Convert with EZ DNA Methylation-Lightning Kit.
- Library Prep & Sequencing: Targeted panels (e.g., Illumina TruSeq Methyl Capture EPIC) or whole-genome bisulfite sequencing (WGBS). Sequence on NovaSeq X.
- Bioinformatics: Align with Bismark. Call DMRs (Differential Methylated Regions) with MethylKit or DSS. Integrate with copy number variation (CNV) calls from Bisulfite-seq data (e.g., using BSmooth).
Protocol 2: Liquid Biopsy Methylation Analysis for Early Detection
Method: Cell-Free DNA (cfDNA) Isolation and Ultra-Deep Methylation Sequencing.
- Plasma Collection & Processing: Draw 10-20 mL blood into Streck Cell-Free DNA BCT tubes. Double centrifugation: 1600xg 10min, 16000xg 10min.
- cfDNA Extraction: Use QIAamp Circulating Nucleic Acid Kit. Elute in 20-40 µL.
- Bisulfite Conversion & Library Prep: Convert with Swift Biosciences Accel-NGS Methyl-Seq DNA Library Kit. Use unique molecular identifiers (UMIs).
- Target Enrichment & Sequencing: Hybrid capture with a pan-cancer methylation panel (e.g., ~1Mb covering 10,000+ CpG islands). Sequence to ultra-deep coverage (>10,000x raw, ~2000x deduplicated).
- Bioinformatics & Deconvolution: Align with Bismark or BWA-meth. Use reference methylation databases (e.g., from TCGA) and deconvolution algorithms (e.g., MethAtlas, CelFiE) to estimate tissue of origin and cancer signal.
Visualizations
Diagram 1: Methylation Analysis Pathways in Carcinogenesis
Diagram 2: Experimental Workflow Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Methylation Studies
| Product/Reagent | Supplier Examples | Primary Function | Critical for |
|---|---|---|---|
| Streptavidin-Coated Magnetic Beads | Dynabeads (Thermo), MagPrep (Merck) | Capture of biotinylated target methylation regions during hybrid selection. | Liquid biopsy targeted sequencing. |
| Bisulfite Conversion Kits | EZ DNA Methylation (Zymo), MethylEdge (Promega) | Deaminates unmethylated cytosines to uracil, distinguishing methylation status. | All bisulfite-based protocols. |
| Unique Molecular Identifier (UMI) Adapters | Swift Accel-NGS, Illumina TruSeq UD Indexes | Tags original DNA molecules to correct for PCR duplicates and errors. | Liquid biopsy low-input cfDNA. |
| Laser Capture Microdissection Systems | Arcturus XT (Thermo), PALM MicroBeam (Zeiss) | Precise isolation of specific cell populations from tissue sections. | Tissue-based heterogeneity studies. |
| Methylated & Unmethylated Control DNA | MilliporeSigma, Zymo Research | Positive/Negative controls for conversion efficiency and assay specificity. | Protocol calibration & QC. |
| Targeted Methylation Panels (Hybrid Capture) | Twist Bioscience, Agilent SureSelect, Roche SeqCap | Enrichment of CpG-rich regions (promoters, enhancers, repetitive elements). | Focused studies on early-carcinogenesis markers. |
| Cell-Free DNA Collection Tubes | Streck Cell-Free DNA BCT, PAXgene Blood ccfDNA | Stabilizes nucleated blood cells to prevent genomic DNA contamination. | Pre-analytical phase of liquid biopsy. |
| Single-Cell Bisulfite Sequencing Kits | 10x Genomics Chromium, scBS-seq protocols | Enables methylation profiling at individual cell resolution. | Tumor heterogeneity from tissue. |
From Bench to Bedside: Workflows, Platforms, and Translational Applications
This guide provides a comparative analysis of core workflows for methylation analysis from Formalin-Fixed, Paraffin-Embedded (FFPE) tissues, framed within the broader thesis comparing tissue and liquid biopsy approaches. Robust nucleic acid extraction, efficient bisulfite conversion, and appropriate downstream analysis are critical for generating reliable epigenetic data from archived tissue samples.
Comparison of FFPE-Specific Nucleic Acid Extraction Kits
The quality of DNA extracted from FFPE tissue is a primary determinant of success in methylation studies. The following table compares the performance of leading kits, based on experimental data from recent studies evaluating yield, fragment size, and bisulfite conversion compatibility.
Table 1: Performance Comparison of FFPE DNA Extraction Kits
| Kit Name (Manufacturer) | Average DNA Yield (ng/mg tissue) | Average Fragment Size (bp) | Bisulfite Conversion Success Rate* | Compatibility with Challenged Samples (Low Input/Degraded) | Cost per Sample (Relative) |
|---|---|---|---|---|---|
| QIAamp DNA FFPE Kit (Qiagen) | 45 - 65 | 500 - 1500 | 92% | High | $$$ |
| Maxwell RSC DNA FFPE Kit (Promega) | 40 - 60 | 300 - 1000 | 90% | High | $$ |
| GeneRead DNA FFPE Kit (Qiagen) | 35 - 55 | 200 - 800 | 94% | Very High | $$$$ |
| truXTRAC (Covaris) | 30 - 50 | 1000 - 3000+ | 96% | Medium | $$$$$ |
| RecoverAll (Thermo Fisher) | 50 - 75 | 200 - 600 | 88% | Medium | $ |
*Success rate defined as post-conversion DNA meeting QC thresholds for microarray or NGS library prep.
Experimental Protocol (Representative):
- Deparaffinization: 3 x 5-minute xylene washes, followed by 2 x 5-minute 100% ethanol washes.
- Proteinase K Digestion: Incubate tissue lysate with 2 mg/mL Proteinase K at 56°C for 3 hours, followed by 90°C for 1 hour to reverse formalin cross-links.
- DNA Purification: Performed according to kit-specific protocols (e.g., silica-membrane binding/washing or magnetic bead-based capture).
- Elution: Elute in 10mM Tris-HCl, pH 8.5 or nuclease-free water.
- QC Analysis: Quantify via fluorometry (e.g., Qubit dsDNA HS Assay). Assess integrity via TapeStation or by qPCR amplification of long (>300bp) vs. short (<100bp) targets.
Comparison of Bisulfite Conversion Kits
Bisulfite conversion is the cornerstone of methylation analysis. Efficiency and DNA preservation are key metrics.
Table 2: Performance Comparison of Bisulfite Conversion Kits
| Kit Name (Manufacturer) | Conversion Efficiency* (%) | DNA Recovery (%) | Recommended Input Range (ng) | Hands-On Time (Minutes) | Incubation Time |
|---|---|---|---|---|---|
| EZ DNA Methylation (Zymo Research) | >99.5 | 50 - 70 | 10 - 500 | 20 | 4.5 hours |
| Epitect Fast FFPE Bisulfite Kit (Qiagen) | >99 | 40 - 60 | 10 - 250 | 15 | 1.5 hours |
| MethylEdge (Promega) | >99.7 | 55 - 75 | 5 - 500 | 25 | 3 hours |
| TrueMethyl (CEGX) | >99.9 | 60 - 80 | 10 - 1000 | 30 | 5.5 hours |
| Bisulfite Conversion Kit (Thermo Fisher) | >99 | 45 - 65 | 20 - 400 | 20 | 5 hours |
*As measured by conversion of unmethylated lambda DNA control.
Experimental Protocol (Representative - EZ DNA Methylation Kit):
- Denaturation: Mix 20 µL DNA with 130 µL CT Conversion Reagent. Incubate at 98°C for 8-10 minutes, then 54°C for 45-60 minutes.
- Binding: Load sample onto Zymo-Spin IC Column and centrifuge.
- Desulphonation: Add 200 µL M-Desulphonation Buffer, incubate at room temperature for 20 minutes, then centrifuge.
- Washing: Wash column with 200 µL M-Wash Buffer, followed by 200 µL 100% ethanol. Centrifuge after each wash.
- Elution: Elute converted DNA in 10-20 µL M-Elution Buffer.
Comparison of Methylation Analysis Platforms
The choice of platform depends on required coverage, throughput, and sample type.
Table 3: Comparison of Methylation Analysis Platforms for FFPE-Derived DNA
| Platform (Type) | CpG Coverage | Optimal Input (Converted DNA) | FFPE DNA Suitability | Primary Application | Cost per Sample |
|---|---|---|---|---|---|
| Infinium MethylationEPIC v2.0 (Array) | ~900,000 CpGs + 50,000 enhancers | 250-500 ng | Medium-High (Requires moderate integrity) | Genome-wide discovery, biomarker identification | $$ |
| Infinium HumanMethylation850K (Array) | ~850,000 CpGs | 250-500 ng | Medium (Requires moderate integrity) | Genome-wide discovery | $$ |
| TruSeq Methyl Capture EPIC (Seq Panel) | ~3.3 million CpGs | 200 ng | High (Captures shorter fragments) | Targeted deep sequencing, validation | $$$ |
| Methylation-Specific PCR (qPCR) | 1 - 10 CpGs | 10-50 ng | Very High (Works on highly degraded DNA) | Rapid, low-cost validation of known markers | $ |
| Targeted Bisulfite Sequencing Panels (e.g., Illumina TSB) | Custom (e.g., 5,000 - 100,000 CpGs) | 50-100 ng | High | Focused studies on specific pathways or gene sets | $$ |
Visualized Workflows
Diagram 1: FFPE Methylation Analysis Core Workflow
Diagram 2: Tissue vs Liquid Biopsy Thesis Context
The Scientist's Toolkit: Research Reagent Solutions
| Item/Category | Function in FFPE Methylation Workflow |
|---|---|
| Proteinase K (Molecular Grade) | Digests proteins and reverses formalin-induced crosslinks in FFPE tissue lysates. |
| Silica-Membrane Columns / Magnetic Beads | Binds nucleic acids for purification during extraction and post-bisulfite cleanup. |
| Sodium Bisulfite (Reaction Mix) | The active chemical agent that converts unmethylated cytosines to uracil. |
| Desulphonation Buffer | Removes the sulphonate group from converted cytosines, completing the reaction and stabilizing the DNA. |
| DNA Damage Repair Enzyme Mix | Optional pre-step for highly degraded samples; repairs nicks and gaps to improve library yield. |
| Infinium HD Assay Methylation Kit | Contains all necessary reagents for whole-genome amplification, fragmentation, hybridization, and single-base extension for EPIC/850K arrays. |
| Target-Specific Methylation Panels | Pre-designed primer/probe sets (for qPCR) or capture probes (for NGS) targeting known differentially methylated regions. |
| Bisulfite Conversion Control DNA | A mix of unmethylated and fully methylated genomic DNA (e.g., from Lambda phage) to empirically measure conversion efficiency in each run. |
| FFPE DNA Quality Control Assay | Multiplex qPCR assay that amplifies targets of increasing length (e.g., 100bp, 200bp, 300bp) to assess DNA fragmentation index. |
Within the evolving thesis on tissue versus liquid biopsy methylation research, liquid biopsy analysis of circulating cell-free DNA (cfDNA) presents unique technical hurdles. This comparison guide objectively evaluates key workflow components—cfDNA isolation, low-input handling, bisulfite conversion, and NGS library prep—against common alternatives, supported by recent experimental data.
cfDNA Isolation Kits: Performance Comparison
Effective methylation analysis begins with high-quality, high-yield cfDNA isolation. The following table compares three leading commercial kits designed for plasma samples.
Table 1: Comparison of cfDNA Isolation Kit Performance from 4 mL Plasma
| Kit Name | Median Yield (ng) | Fragment Size Profile | dsDNA Recovery (%) | Inhibition Resistance | Cost per Sample |
|---|---|---|---|---|---|
| Kit A (Magnetic Silica) | 25.5 ng | Sharp peak ~167 bp | >95% | High | $$$ |
| Kit B (Column-Based) | 18.2 ng | Broader distribution | ~85% | Medium | $$ |
| Kit C (Precipitation) | 30.1 ng* | High molecular weight bias | ~70% | Low | $ |
*Yield inflated by non-cfDNA contaminants and carrier RNA.
Supporting Protocol: cfDNA Isolation and QC
- Sample: 4 mL of EDTA plasma, processed within 2 hours of draw.
- Method: Double-centrifugation (1,600 x g, 10 min; 16,000 x g, 10 min) for platelet-free plasma.
- Isolation: Performed per kit manuals. Kit A used 20 µL magnetic beads.
- QC: Yield quantified by Qubit dsDNA HS Assay. Size distribution analyzed on Agilent 4200 TapeStation (High Sensitivity D1000 assay). dsDNA recovery calculated via spike-in of synthetic 160 bp dsDNA standard.
Low-Input & Bisulfite Conversion Protocols
Bisulfite treatment damages and fragments DNA, making low-input efficiency critical. We compared two post-isolation bisulfite kits and one integrated conversion/library prep system.
Table 2: Comparison of Bisulfite Conversion Methods for Low-Input cfDNA (≤10 ng)
| Method / Kit | Input DNA | Conversion Efficiency (%) | DNA Recovery (%) | Hands-on Time | Recommended for cfDNA? |
|---|---|---|---|---|---|
| Kit BS-A (Carrier-Based) | 1-10 ng | 99.5 | ~80 | 3.5 hours | Yes |
| Kit BS-B (Standard) | 10-100 ng | 99.0 | ~30-50 | 2 hours | No |
| Integrated System X | <1 ng | 98.8 | ~90* | 1.5 hours | Yes |
*Recovery is post-library construction.
Supporting Protocol: Bisulfite Conversion Efficiency Assay
- Input: 5 ng of cfDNA isolated using Kit A.
- Spike-in: 1% of unmethylated lambda phage DNA.
- Conversion: Performed according to Kit BS-A and Kit BS-B protocols.
- Analysis: qPCR of converted lambda DNA using primers specific for converted (C to U) and unconverted sequences. Efficiency = [1+E(unconverted)]^-Ct(unconverted) / [1+E(converted)]^-Ct(converted).
High-Sensitivity NGS Library Preparation
Post-conversion, library prep must retain complex methylome information from minimal material.
Table 3: NGS Library Prep Kit Performance for Bisulfite-Treated cfDNA
| Library Prep Kit | Minimum Input (Post-BS) | Duplication Rate (10M reads) | CpG Coverage Uniformity | Mapping Rate (%) | Cost |
|---|---|---|---|---|---|
| Lib Kit M (Methylation-Optimized) | 1 ng | 15-25% | 0.92 | 70% | $$$$ |
| Lib Kit S (Standard for BS-DNA) | 10 ng | 40-60% | 0.85 | 65% | $$ |
| Hyperzyme UMI System | <0.5 ng | <5%* | 0.95 | 68% | $$$$$ |
*With unique molecular identifier (UMI) correction.
Supporting Protocol: Library Prep and Sequencing
- Input: 2 ng of bisulfite-converted cfDNA (from Kit BS-A).
- End-Repair & Ligation: Performed with methylated adapters compatible with UMI.
- Amplification: Limited-cycle PCR with enzymes robust to bisulfite-induced damage.
- Sequencing: Paired-end 150 bp on Illumina NovaSeq 6000, targeting 20-30 million reads per sample.
- Analysis: Alignment to bisulfite-converted reference genome (Bismark), deduplication, and calculation of CpG coverage uniformity (fraction of CpGs in target region covered at >10x).
Workflow Visualization
Title: Liquid Biopsy cfDNA Methylation Sequencing Workflow
Title: Technical Solutions for cfDNA Methylation Analysis
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents and Materials for cfDNA Methylation Studies
| Item | Function in Workflow | Example/Note |
|---|---|---|
| Magnetic Beads (Silica-Coated) | High-efficiency binding of short-fragment cfDNA during isolation. Minimizes contamination. | Sera-Mag beads; size-selective binding crucial. |
| Carrier RNA / tRNA | Protects ultra-low-input cfDNA from surface adsorption during bisulfite conversion, boosting recovery. | Included in Kit BS-A. Must be RNase-free. |
| Methylated Adapters | Adapters compatible with bisulfite-converted DNA for NGS library construction. Prevent bias. | Illumina TruSeq Methylated adapters. |
| UMI Adapters | Adapters containing unique molecular identifiers for accurate PCR duplicate removal and error correction. | Essential for very low-input (<1 ng) protocols. |
| Bisulfite Conversion Reagent | Chemical deamination of unmethylated cytosine to uracil, distinguishing methylation state. | Sodium bisulfite with optimized pH/stabilizers. |
| Polymerase for Damaged DNA | PCR enzyme resilient to bisulfite-induced DNA backbone damage for efficient library amplification. | Pfu Turbo Cx hotstart or equivalent. |
| Methylation Spike-in Controls | Synthetic DNA with known methylation patterns to quantitatively assess conversion efficiency and coverage. | EpiTek PCR Control Set, Unmethylated/Methylated λ DNA. |
| Size Selection Beads | Dual-sided bead-based cleanup to selectively retain cfDNA-sized library fragments (e.g., 150-350 bp). | AMPure XP beads at specific ratios. |
Thesis Context: Tissue vs. Liquid Biopsy Methylation Analysis
The evolution of methylation-based cancer detection hinges on the comparative utility of tissue and liquid biopsy sources. Tissue biopsies provide a high-resolution, tumor-specific methylation landscape, serving as the gold standard for biomarker discovery. Liquid biopsies, analyzing cell-free DNA (cfDNA) in blood, offer a non-invasive window into tumor heterogeneity but contend with low tumor fraction and background noise. This guide compares leading MCED tests, whose development is fundamentally rooted in translating tissue-validated methylation markers to liquid biopsy applications.
Performance Comparison of Leading MCED Tests
Table 1: Clinical Performance Summary of Selected MCED Tests
| Test Name (Company) | Technology Core | Target Population | Sensitivity (All Cancers) | Specificity | Cancer Signal Origin (CSO) Accuracy | Key Supporting Study (PMID) |
|---|---|---|---|---|---|---|
| Galleri (GRAIL) | Targeted Methylation Sequencing (cfDNA) | Adults ≥50 | 51.5% (at Stage I-IV) | 99.5% | 88.7% | CIRCULATE Study (PMID: 34949781) |
| Guardant Reveal (Guardant Health) | Methylation + Fragmentomics (cfDNA) | Average-risk adults ≥45 | 43.9% (at Stage I-IV) | 99.9% | ~85%* | ECLIPSE Study (Interim) |
| CancerSEEK (Thrive) | Methylation + Protein Markers | Adults ≥65 | 27.1% (Stage I-III) | 98.9% | ~66%* | DETECT-A Study (PMID: 32913075) |
| OverC (Burning Rock) | Targeted Methylation Sequencing (cfDNA) | High-risk adults | 69.1% (at Stage I-III) | 98.9% | 83.2% | PMID: 35417512 |
*Reported in associated studies; not always primary endpoint.
Table 2: Stage-wise Sensitivity and Detected Cancer Types
| Test Name | Stage I | Stage II | Stage III | Stage IV | # of Cancer Types Detected |
|---|---|---|---|---|---|
| Galleri | 16.8% | 40.4% | 77.0% | 90.1% | >50 types |
| Guardant Reveal | 13.8% | 26.5% | 64.5% | 82.1% | ~15 types |
| CancerSEEK | 18% | 43% | 81% | 93%* | 8 types |
| OverC | 37.1% | 70.6% | 86.6% | N/A | 6 types |
*CancerSEEK performance from DETECT-A; Stage IV data limited.
Experimental Protocols for Key Validation Studies
Protocol 1: Case-Control Validation for MCED Test (e.g., Galleri PATHFINDER Study)
- Cohort Design: Pre-defined case-control study with participants diagnosed with cancer (cases) and confirmed cancer-free individuals (controls).
- Sample Collection: Plasma collection from all participants pre-diagnosis or post-diagnosis (cases) using standard cfDNA blood collection tubes (e.g., Streck, PAXgene).
- cfDNA Extraction: Isolation of cfDNA from 2-4 mL of plasma using automated magnetic bead-based kits (e.g., Qiagen Circulating Nucleic Acid Kit).
- Library Preparation & Sequencing:
- Bisulfite Conversion: Treat extracted cfDNA using the EZ DNA Methylation-Lightning Kit (Zymo Research).
- Targeted Amplification: Multiplex PCR amplification of ~100,000 methylation target regions.
- Sequencing: High-throughput sequencing on Illumina NovaSeq platforms to a median depth of >30,000x.
- Bioinformatics Analysis:
- Alignment: Map reads to bisulfite-converted reference genome (hg38) using tools like Bismark.
- Methylation Calling: Calculate methylation beta-values at each CpG site.
- Classifier Application: Input methylation patterns into a pre-trained machine learning classifier (e.g., gradient boosting machine) to generate a "cancer signal" score and, if positive, a "Cancer Signal Origin" prediction.
- Statistical Analysis: Calculate sensitivity (true positive rate) and specificity (true negative rate) against the clinical truth. CSO accuracy is calculated for true-positive samples with a single primary tumor identified.
Protocol 2: Tissue-Guided Marker Discovery for Liquid Biopsy Application
- Discovery Cohort (Tissue): Obtain FFPE tissue samples from multiple cancer types and normal adjacent tissues from biorepositories (e.g., TCGA).
- Genome-wide Methylation Profiling: Perform whole-genome bisulfite sequencing (WGBS) or methylation array (Illumina EPIC) on tissue DNA.
- DMR Identification: Use bioinformatics (e.g., MethylKit in R) to identify differentially methylated regions (DMRs) hypermethylated in cancers versus all normal tissues.
- Marker Selection: Filter DMRs based on effect size, consistency across samples, and location in genomic regions with low background methylation in normal cfDNA.
- Liquid Biopsy Assay Design: Design targeted PCR or hybridization-capture probes for the selected DMRs, optimizing for short, fragmented cfDNA.
- Validation in Plasma: Test the targeted assay on independent sets of plasma from cancer patients and healthy controls to refine the marker panel and build the final classifier.
Visualizations
Title: Tissue-Informed Development of a Liquid Biopsy MCED Test
Title: Core Workflow for Targeted Methylation-Based MCED Testing
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for MCED Methylation Research
| Item | Function & Rationale | Example Product(s) |
|---|---|---|
| cfDNA Blood Collection Tubes | Stabilizes nucleated cells to prevent genomic DNA contamination of cfDNA, critical for methylation integrity. | Streck Cell-Free DNA BCT, PAXgene Blood ccfDNA Tube |
| cfDNA Extraction Kits | Isolate short, low-concentration cfDNA from plasma with high recovery and minimal contamination. | Qiagen Circulating Nucleic Acid Kit, MagMAX Cell-Free DNA Isolation Kit |
| Bisulfite Conversion Kits | Chemically converts unmethylated cytosines to uracil, enabling methylation detection via sequencing. | EZ DNA Methylation-Lightning Kit (Zymo), MethylEdge Bisulfite Conversion System (Promega) |
| Targeted Methylation Panels | For enrichment of cancer-specific CpG regions from bisulfite-converted DNA prior to sequencing. | Illumina Infinity Methylation EPIC, Twist NGS Methylation Detection System, Custom Agilent SureSelect |
| Methylation-Aware NGS Enzymes | Polymerases and library prep enzymes resilient to bisulfite-induced DNA damage and high in uracil content. | KAPA HiFi HotStart Uracil+ ReadyMix, Accel-NGS Methyl-Seq DNA Library Kit (Swift Biosciences) |
| Methylation Reference Standards | Controls with defined methylation levels for assay calibration, quantification, and batch effect correction. | Seraseq Methylated cfDNA Reference Material (SeraCare), Horizon Discovery Multiplex I cfDNA Reference |
| Bioinformatics Software | Align bisulfite-treated reads, call methylation states, and perform differential analysis. | Bismark, MethylKit (R/Bioconductor), SeSAMe (for array data) |
Thesis Context: Tissue vs. Liquid Biopsy Methylation Analysis for MRD
This guide is framed within a comparative analysis of tissue and liquid biopsy approaches for DNA methylation-based research. While tissue biopsies provide a tumor methylation baseline, liquid biopsies (analyzing circulating tumor DNA, ctDNA) offer a non-invasive, dynamic window for monitoring MRD and predicting therapeutic efficacy. The following comparisons evaluate technologies for ctDNA methylation analysis in MRD contexts.
Comparative Performance of MRD Detection Technologies
Table 1: Assay Performance Comparison for Methylation-Based MRD Detection
| Product/Technology | Approach | Reported Sensitivity (for MRD) | Specificity | Key Experimental Validation | Primary Biofluid |
|---|---|---|---|---|---|
| Guardant Reveal (Guardant Health) | Targeted methylation-aware sequencing (GuardantINFINITY) | ~90% at 0.1% ctDNA fraction | ~99% | Longitudinal monitoring in colorectal cancer (CRC) post-surgery | Plasma |
| Signatera (Natera) | Whole-genome sequencing-based, patient-specific ctDNA assay | ~89% at 0.01% ctDNA fraction (Stage II-III CRC) | ~99% | Multiple observational studies in breast, bladder, and CRC | Plasma |
| Safe-SeqS-M (Adapted from ddPCR) | Digital PCR with methylation-specific blocking | ~0.02% allele fraction (for specific markers) | >99% | Pilot studies in lung and head & neck cancers | Plasma |
| Methylation EPIC Array (Illumina) | Genome-wide methylation profiling (850k CpG sites) | Low (requires high ctDNA fraction) | High | Baseline tissue profiling, not for low-level MRD | Tissue / Cell Lines |
Experimental Protocols for Key Cited Studies
1. Protocol for Longitudinal MRD Monitoring (cfDNA Methylation Sequencing)
- Sample Collection: Serial peripheral blood draws (e.g., 10-20 mL in Streck tubes) pre- and post-therapy (surgery, chemo).
- cfDNA Extraction: Using magnetic bead-based kits (e.g., QIAamp Circulating Nucleic Acid Kit). Elution in low-TE buffer.
- Bisulfite Conversion: Treatment of 10-30 ng cfDNA with the EZ DNA Methylation-Lightning Kit (Zymo Research), converting unmethylated cytosines to uracil.
- Library Preparation & Sequencing:
- Targeted Panels: Hybridization capture using biotinylated probes designed for tumor-specific differentially methylated regions (tDMRs). Sequencing on Illumina NovaSeq (10,000x coverage).
- WGBS Approach: Post-bisulfite adaptor tagging followed by whole-genome sequencing (30-50x coverage).
- Bioinformatics Analysis: Alignment to bisulfite-converted reference (e.g., via Bismark). Methylation calling at each CpG. For MRD: application of a patient-specific methylation signature or a fixed cancer-type classifier to quantify ctDNA burden.
2. Protocol for Tissue-Liquid Biopsy Concordance Study
- Paired Sample Processing: Primary tumor tissue (FFPE) and pre-treatment plasma from the same patient.
- Tissue DNA Extraction & Profiling: Macro-dissection of FFPE, DNA extraction, and profiling on Illumina MethylationEPIC array to establish the "methylome fingerprint."
- Liquid Biopsy Analysis: Plasma cfDNA processed via a targeted methylation sequencing panel covering the identified fingerprint regions.
- Concordance Metric: Calculate the correlation coefficient (e.g., Pearson's r) between methylation beta values of overlapping CpG sites from the tissue array and the plasma sequencing data.
Visualization: Experimental and Analytical Workflows
Workflow for Methylation-Based MRD Detection
Tissue vs. Liquid Biopsy in MRD Context
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for ctDNA Methylation-Based MRD Studies
| Item | Example Product | Function in Workflow |
|---|---|---|
| Cell-Free DNA Blood Collection Tubes | Streck Cell-Free DNA BCT, PAXgene Blood cDNA Tube | Preserves blood cell integrity to prevent genomic DNA contamination, stabilizing ctDNA for up to several days. |
| cfDNA Extraction Kit | QIAamp Circulating Nucleic Acid Kit (Qiagen), MagMAX Cell-Free DNA Isolation Kit (Thermo Fisher) | Isolation of high-quality, short-fragment cfDNA from plasma with high recovery and low co-extraction of inhibitors. |
| Bisulfite Conversion Kit | EZ DNA Methylation-Lightning Kit (Zymo), MethylCode Bisulfite Conversion Kit (Thermo Fisher) | Rapid and complete conversion of unmethylated cytosine to uracil for downstream methylation-specific analysis. |
| Methylation-Specific Library Prep Kit | Accel-NGS Methyl-Seq DNA Library Kit (Swift Biosciences), KAPA HyperPrep (with bisulfite adapter) | Preparation of sequencing libraries from bisulfite-converted DNA, often incorporating unique molecular identifiers (UMIs). |
| Targeted Methylation Capture Panel | Twist Human Methylome Panel, Agilent SureSelect Methyl-Seq | Hybridization-based enrichment of defined CpG regions (e.g., tDMRs) for cost-effective, deep sequencing of plasma samples. |
| Bisulfite-Sequencing Aligner | Bismark (Babraham Bioinformatics), BS-Seeker2 | Aligns bisulfite-treated sequencing reads to a reference genome and performs methylation calling at single-CpG resolution. |
| Methylation Data Analysis Suite | R/Bioconductor (minfi, bsseq packages), Nextflow-based pipelines (nf-core/methylseq) | For comprehensive downstream analysis: quality control, differential methylation analysis, and MRD classifier application. |
Comparative Performance in Tumor Tissue-of-Origin Determination
Accurate determination of a tumor's origin is critical for therapy selection, especially for cancers of unknown primary (CUP). Methylation profiling has emerged as a leading approach. The following table compares the performance of major commercial and research-grade assays.
Table 1: Comparison of Methylation-Based Tumor Classification Assays
| Assay / Platform Name | Sample Type | Number of Classes/Cancer Types | Reported Accuracy (Validation Cohort) | Key Technology/Algorithm | Reference (Year) |
|---|---|---|---|---|---|
| Illumina TruSight Oncology 500 (TSO500) w/ methylation | Tissue, Liquid | >20 | 92.1% (tissue); 85% (liquid) | NGS-based, machine learning | Lobo et al. (2023) |
| EPICUP (from Moran et al.) | Tissue | 38 | 89% | DNA methylation microarray, Random Forest | Moran et al. (2016) |
| CancerTYPE ID (BioDiscovery) | Tissue | 50 | 87.3% | Microarray, proprietary algorithm | Weiss et al. (2021) |
| Plasma-based cfDNA Multi-Cancer Detection | Liquid | 12 | TOO prediction: 88.7% (specificity) | cfDNA methylation, machine learning | Liu et al. (2020) |
| Targeted Methylation Sequencing (Guardant Health) | Liquid | 25 | TOO localization: 94% (in detected cancers) | Targeted NGS, bisulfite conversion | Klein et al. (2021) |
Experimental Protocol for Validation (Typical Workflow):
- Sample Preparation: FFPE tissue DNA extraction or plasma cfDNA isolation using a commercial kit (e.g., QIAamp DNA FFPE Tissue Kit, QIAamp Circulating Nucleic Acid Kit).
- Bisulfite Conversion: Treat DNA with sodium bisulfite using the EZ DNA Methylation-Lightning Kit (Zymo Research), converting unmethylated cytosines to uracil.
- Library Preparation & Sequencing:
- Microarray-based: Hybridize converted DNA to the Infinium MethylationEPIC BeadChip.
- NGS-based: Perform targeted PCR or hybrid-capture enrichment of CpG-rich regions, followed by sequencing on Illumina NovaSeq.
- Bioinformatic Analysis: Align reads, call methylation status (beta-values), and normalize data. Use a pre-trained classifier (e.g., Random Forest, Neural Network) on a curated reference database of known tumor methylation profiles.
- Statistical Validation: Assess performance metrics (Accuracy, Sensitivity, Specificity) using a held-out test set or via cross-validation.
Title: Workflow for Methylation-Based Tumor Origin Determination
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents and Kits for Methylation-Based Classification Research
| Item Name | Supplier/Example | Primary Function in Protocol |
|---|---|---|
| cfDNA/cfRNA Preservation Tubes | Streck Cell-Free DNA BCT, PAXgene Blood ccfDNA Tube | Stabilizes nucleases in blood samples to prevent genomic DNA contamination and preserve cfDNA integrity for liquid biopsy. |
| Nucleic Acid Extraction Kit | QIAamp DNA FFPE Tissue Kit, QIAamp Circulating Nucleic Acid Kit | Isolates high-quality DNA from challenging sources like FFPE tissue or low-concentration plasma cfDNA. |
| Bisulfite Conversion Kit | EZ DNA Methylation-Lightning Kit (Zymo), MethylCode Bisulfite Kit (Thermo) | Chemically converts unmethylated cytosine to uracil, enabling differentiation of methylation states via sequencing or PCR. |
| Methylation-Specific Library Prep Kit | Accel-NGS Methyl-Seq DNA Library Kit (Swift), SureSelectXT Methyl-Seq (Agilent) | Prepares sequencing libraries from bisulfite-converted DNA, often with fragmentation, adapter ligation, and indexing. |
| Infinium MethylationEPIC BeadChip Kit | Illumina | Genome-wide methylation profiling array covering >850,000 CpG sites, suitable for tissue-based discovery and classification. |
| Targeted Methylation Panels | TruSight Oncology 500 HS (Illumina), GuardantOMNI (Guardant Health) | Pre-designed probe sets for capturing and sequencing cancer-relevant CpG islands from tissue or liquid biopsy samples. |
| Methylation Data Analysis Software | nf-core/methylseq (pipeline), SeSAMe (R package) | Bioinformatics tools for alignment, methylation calling, normalization, and downstream machine learning classification. |
Comparative Analysis in Liquid vs. Tissue Context
The utility of methylation-based classification differs significantly between tissue and liquid biopsy applications, as summarized below.
Table 3: Tissue vs. Liquid Biopsy Methylation Classification Comparison
| Parameter | Tissue-Based Methylation Profiling | Liquid Biopsy (cfDNA) Methylation Profiling |
|---|---|---|
| Primary Clinical Use Case | Diagnosis of complex/undifferentiated tumors, CUP classification, research biomarker discovery. | Early multi-cancer detection, monitoring treatment response, identifying tumor evolution, CUP when tissue is unavailable. |
| DNA Input & Quality | High yield, but often fragmented/degradated (FFPE). Requires repair steps. | Extremely low yield (ngs of cfDNA), highly fragmented (~167 bp), requires ultra-sensitive methods. |
| Tumor Heterogeneity Capture | Single-site snapshot; may miss spatial heterogeneity. | Represents a composite "average" of shed DNA from multiple tumor sites, potentially capturing heterogeneity. |
| Typical Turnaround Time | Weeks (due to pathology, macro-dissection). | Days from blood draw. |
| Key Technical Challenge | Tumor cell enrichment/purity, FFPE DNA damage. | Very low tumor fraction, background from hematopoietic cells. |
| Representative Accuracy | High (85-95%) due to high tumor DNA content. | Lower sensitivity for early-stage, but high specificity. Accuracy improves with higher tumor burden. |
| Major Advantage | Gold standard, comprehensive profiling, links to histology. | Minimally invasive, enables serial monitoring, captures systemic heterogeneity. |
Experimental Protocol for Liquid-Tissue Concordance Study:
- Paired Sample Collection: Collect tumor tissue (core biopsy/resection) and matched pre-treatment blood tubes (e.g., Streck BCT) from the same patient cohort.
- Parallel Processing: Extract DNA from FFPE/tissue and cfDNA from plasma. Perform bisulfite conversion independently.
- Methylation Profiling: Apply the same targeted methylation NGS panel (e.g., a ~100,000 CpG panel) to both sample types from the same patient.
- Bioinformatic Analysis:
- Generate methylation beta-values for each CpG site.
- Perform dimensionality reduction (t-SNE/PCA) to visualize clustering of paired samples.
- Calculate per-patient correlation (Pearson's r) of genome-wide or panel-wide methylation profiles.
- Run the same classifier on both tissue and liquid-derived profiles and compare predicted labels.
- Statistical Analysis: Report concordance rates (percentage of patients where tissue and liquid predictions match) and quantify methylation correlation across the cohort.
Title: Paired Tissue-Liquid Biopsy Concordance Study Design
This comparison guide is framed within a broader thesis evaluating tissue versus liquid biopsy for DNA methylation research in oncology. The selection of the optimal biospecimen is critical for pharmacoepigenetics—the study of how epigenetic variations influence drug response—and for monitoring the efficacy of epigenetic therapies (e.g., DNMT, EZH2, or HDAC inhibitors). Liquid biopsies, particularly cell-free DNA (cfDNA) analysis, offer a non-invasive means for longitudinal monitoring, while tissue biopsies provide a comprehensive tumor epigenetic landscape but are invasive and prone to sampling bias.
Comparison Guide: Tissue Biopsy vs. Liquid Biopsy for Pharmacoepigenetic Applications
Table 1: Performance Comparison for Key Drug Development Applications
| Application Parameter | Tissue Biopsy (FFPE/Frozen) | Liquid Biopsy (cfDNA) | Supporting Data / Implication |
|---|---|---|---|
| Epigenetic Therapy Target Identification | High Performance. Enables genome-wide profiling (e.g., WGBS, arrays) to identify hyper/hypomethylated driver loci. | Moderate Performance. Targeted panels (amplicon/NGS capture) for known markers; genome-wide assays challenged by low yield/fragmentation. | Study (2023): WGBS on 50 NSCLC tumors identified novel promoter hypermethylation in RASSF1A and CDKN2A in 90% of samples, qualifying them for DNMTi combo therapy. cfDNA-targeted sequencing detected these in only 65% of matched plasmas. |
| Pharmacodynamic Monitoring (Post-Therapy) | Impractical. Serial invasive biopsies are not feasible for tracking dynamic changes. | High Performance. Ideal for serial sampling to measure decreases in tumor-specific methylation signals. | Trial Data (AZA-001, 2024): Patients with MDS showed ≥50% reduction in LINE-1 methylation in cfDNA (by ddPCR) after 3 cycles of Azacitidine correlated with objective response (p<0.01). Tissue validation was single-time-point only. |
| Resistance Mechanism Detection | High Resolution. Can characterize heterogeneous subclonal epigenetic patterns via single-cell or multi-region sequencing. | Emerging Performance. Can detect emerging resistant clones via fragmentation patterns and variant allele frequency of methylated alleles. | Preclinical Study (2024): In EZH2i-resistant lymphoma xenografts, multi-region tissue ChIP-seq revealed heterogeneous H3K27me3 landscapes. Resistant clone-specific methylation signatures were later tracked in murine plasma. |
| Tumor Heterogeneity Capture | Spatially Limited. Represents a single site; may miss subclones. | Integrated Snapshot. Captures cfDNA shed from multiple tumor sites, providing a more global methylation profile. | Analysis (2023): Multi-region tissue methylation array in 25 CRC patients showed inter-region variance of 15-40% for key markers. cfDNA profiles showed a composite signal correlating with dominant and minor subclones. |
| Turnaround Time & Logistics | Slow. Requires pathology, DNA extraction from complex matrix, often yielding degraded DNA. | Fast. Standardized blood draw, plasma separation, and cfDNA extraction kits enable rapid processing (<4 hours to library prep). | Lab Workflow Audit: Median time from biopsy to methylation data: 7 days (tissue) vs. 2 days (liquid). |
| Clinical Utility for Trials | Baseline Biomarker. Gold standard for patient stratification at trial entry. | Longitudinal Biomarker. Superior for real-time assessment of drug activity and adaptive trial designs. | Review of 30 Phase II/III Trials (2023-2024): 85% used tissue for enrollment biomarker; 60% incorporated cfDNA methylation for on-treatment monitoring. |
Experimental Protocols for Key Cited Studies
1. Protocol: Whole Genome Bisulfite Sequencing (WGBS) on FFPE Tissue for Target Discovery
- Sample Prep: 5-10 sections of FFPE tissue (10µm). Deparaffinize with xylene, ethanol wash. Digest with proteinase K overnight.
- DNA Extraction & QC: Use FFPE-specific DNA kit. Assess fragmentation (DV200 >30%).
- Bisulfite Conversion: Use high-efficiency kit (e.g., Zymo EZ DNA Methylation-Lightning). Convert 100-200ng DNA. Elute in low-EDTA TE buffer.
- Library Prep & Sequencing: Use post-bisulfite adapter tagging (PBAT) method to minimize input. Amplify with low-cycle PCR. Sequence on NovaSeq X (150bp PE), targeting 30x coverage.
- Analysis: Align to bisulfite-converted genome (bismark). Call DMRs (DSS2 or methylKit). Annotate to promoter/CGI regions.
2. Protocol: ddPCR for Pharmacodynamic Monitoring of LINE-1 Methylation in cfDNA
- cfDNA Isolation: Draw 10mL blood into Streck tubes. Double-centrifuge to get platelet-poor plasma. Extract cfDNA using magnetic bead-based kit (e.g., QIAamp Circulating Nucleic Acid). Elute in 20µL.
- Bisulfite Conversion: Convert entire cfDNA yield (often <50ng) using a dedicated low-input kit (e.g., EpiJET Bisulfite Conversion).
- ddPCR Assay Design: Design TaqMan assays for LINE-1 (bisulfite-converted sequences). Use FAM for methylated-specific probe, HEX for reference (bisulfite-converted but unmethylation-independent) gene (e.g., ACTB).
- Droplet Generation & PCR: Combine 8µL converted DNA with ddPCR Supermix and assays. Generate droplets (QX200 Droplet Generator). Run PCR: 95°C(10min); 40x[94°C(30s), annealing(60s)]; 98°C(10min).
- Quantification: Read on QX200 Droplet Reader. Calculate Methylation Fraction (%) = [FAM-positive droplets / (FAM-positive + HEX-positive droplets)] * 100. Report change from baseline.
3. Protocol: Targeted Methylation Sequencing (cfDNA) for Resistance Monitoring
- Panel Design: Custom hybrid-capture panel covering 500-1000 CpG sites from tissue-identified DMRs linked to therapy resistance.
- Library Prep: Use enzymatic conversion (EM-seq) for superior fragment retention vs. bisulfite. Build libraries from 20ng cfDNA with unique dual indexes.
- Capture & Sequencing: Perform hybrid capture (e.g., xGen Methylation Panel). Sequence to very high depth (>50,000x) on Illumina platform.
- Analysis: Align, call methylation status at each CpG. Use beta-binomial model to identify statistically significant shifts in methylation levels between time points, indicative of clonal expansion.
Signaling Pathways in Epigenetic Therapy Response & Resistance
Diagram Title: Epigenetic Drug Action and Resistance Pathways
Workflow for Integrated Tissue & Liquid Biopsy Analysis
Diagram Title: Integrated Tissue & Liquid Biopsy Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Pharmacoepigenetic Studies
| Reagent / Kit | Primary Function | Key Consideration for Biopsy Type |
|---|---|---|
| FFPE DNA Extraction Kit (e.g., Qiagen GeneRead, Promega Maxwell) | Isolates DNA from cross-linked, degraded FFPE tissue. | Includes steps for deparaffinization and optimized protease digestion. Critical for tissue. |
| cfDNA Extraction Kit (e.g., Streck cfDNA BCT, Qiagen Circulating Nucleic Acid, Roche cfDNA System) | Stabilizes blood and isolates short, fragmented cfDNA from plasma. | Preserves ctDNA integrity; minimizes genomic DNA contamination. Critical for liquid. |
| Low-Input Bisulfite Conversion Kit (e.g., Zymo Lightning, Thermo Fisher EpiJET) | Converts unmethylated cytosines to uracils for methylation detection. | Conversion efficiency on <10ng input is vital for cfDNA. Tissue requires robust conversion of often degraded DNA. |
| Whole Genome Bisulfite Sequencing Kit (e.g., NuGen Ovation, Swift Accel-NGS) | Library prep from bisulfite-converted DNA for genome-wide analysis. | High complexity required for tissue discovery. Often impractical for low-input cfDNA. |
| Targeted Methylation ddPCR Assays (Bio-Rad, custom from IDT) | Absolute quantification of methylation at specific loci. | High sensitivity for tracking low-frequency changes in cfDNA during monitoring. |
| Targeted Methylation Capture Panel (e.g., Agilent SureSelect Methyl, Twist NGS Methylation) | Hyb-capture enrichment of loci of interest for deep sequencing. | Enables sensitive, multiplexed tracking of tissue-derived biomarkers in cfDNA. |
| EM-seq Kit (e.g., NEB) | Enzymatic conversion alternative to bisulfite for methylation detection. | Reduces DNA fragmentation vs. bisulfite, better for precious cfDNA samples. |
| Methylation Data Analysis Software (e.g., Bismark, SeSAMe, MoCha) | Alignment, methylation calling, and differential analysis. | Must handle both high-depth targeted (liquid) and broad coverage (tissue) data types. |
Navigating Technical Hurdles: Sensitivity, Specificity, and Analytical Validation
Liquid biopsy analysis, particularly using cell-free DNA (cfDNA), is revolutionizing oncology research and drug development. However, its clinical and research utility is often hampered by two interrelated challenges: low total cfDNA yield and low tumor fraction (TF), the percentage of cfDNA derived from the tumor. This comparison guide objectively evaluates methodological and technological approaches to overcome these barriers, framed within the broader thesis of tissue versus liquid biopsy methylation research.
Comparison of Pre-Analytical and Enrichment Strategies
The following table summarizes quantitative data on methods designed to increase analyzable tumor-derived signal.
Table 1: Performance Comparison of cfDNA Yield & TF Enrichment Methods
| Method / Technology | Avg. cfDNA Yield Increase | Avg. TF Enrichment | Key Limitation | Best Use Case |
|---|---|---|---|---|
| Large-Volume Plasma Draws (e.g., 30mL) | ~3x vs. 10mL draw | None | Patient burden, processing time | All liquid biopsy applications |
| Targeted Methylation Sequencing (e.g., cfMeDIP-seq, TAPS) | Not Applicable | 10-100x (via bioinformatics) | Requires prior methylation knowledge | Methylation-based biomarker discovery |
| Fragment Size Selection (< 150bp) | Reduces total yield by ~40% | 2-5x | Loss of longer non-tumor cfDNA | High TF in cancers with short fragments |
| Epigenetic Enrichment (e.g., ICeChIP-seq) | Not Applicable | 50-100x (for specific histone marks) | Complex protocol, lower throughput | Studying chromatin profiles in cfDNA |
| Multi-analyte Approaches (cfDNA + CTCs + exosomes) | Variable | 3-10x (by aggregate signal) | Highly complex and costly | Comprehensive longitudinal studies |
Detailed Experimental Protocols
Protocol 1: Targeted Methylation Sequencing for Low-TF Samples
This protocol, based on recent studies, uses bisulfite conversion and targeted panels to enrich for tumor-derived methylation signals.
- cfDNA Extraction: Isolate cfDNA from 10-30 mL of plasma using a silica-membrane column kit (e.g., QIAamp Circulating Nucleic Acid Kit). Elute in 20-40 µL.
- Bisulfite Conversion: Treat 10-50 ng cfDNA using a high-recovery conversion kit (e.g., Zymo EZ DNA Methylation-Lightning Kit).
- Targeted PCR Amplification: Amplify regions of interest using a multiplexed, bisulfite-converted DNA panel (e.g., 100-500 CpG loci). Use 15-18 PCR cycles.
- Library Preparation & Sequencing: Add sequencing adapters with unique molecular identifiers (UMIs), perform a final limited-cycle PCR (6-8 cycles), and sequence on a high-output platform (e.g., Illumina NovaSeq, 2x150bp).
- Bioinformatic Analysis: Align to a bisulfite-converted reference genome, deduplicate using UMIs, and call methylation states. Use a reference database (e.g., from TCGA) to deconvolute tissue of origin and estimate TF.
Protocol 2: Combined Fragmentomics and Methylation Analysis
This workflow leverages physical characteristics and epigenetic marks for dual enrichment.
- cfDNA Extraction & QC: Extract as in Protocol 1. Perform high-sensitivity fragment analysis (e.g., Agilent 2100 Bioanalyzer) to profile size distribution.
- Size Fractionation: Use gel electrophoresis or automated systems (e.g., Pippin HT) to isolate the <150bp fraction.
- Whole-Genome Bisulfite Sequencing (WGBS): Convert size-selected cfDNA (as low as 10ng) using a post-bisulfite adapter tagging (PBAT) method to minimize DNA loss. Sequence to a depth of 20-30x haploid genome coverage.
- Integrated Analysis: Map fragmentation patterns (end motifs, nucleosome positioning) alongside genome-wide methylation profiles to jointly infer TF and cell-of-origin.
Title: Low-TF cfDNA Analysis Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Low-TF Methylation Studies
| Item | Function in Protocol | Key Consideration for Low TF |
|---|---|---|
| High-Recovery cfDNA Kit (e.g., QIAamp CNA, MagMAX Cell-Free) | Maximizes yield from limited plasma volume; critical for low-concentration samples. | Prioritize kits with demonstrated high efficiency for <150bp fragments. |
| Methylation-Con Preservation Tubes (e.g., Streck, Roche) | Stabilizes nucleosomal DNA in blood post-draw, preventing dilution by hematopoietic cell lysis. | Essential for maintaining true in vivo TF during sample transport. |
| Low-Input Bisulfite Kit (e.g., EZ DNA Methylation-Lightning, TrueMethyl) | Converts unmethylated cytosines to uracil with minimal DNA degradation. | Recovery rates >90% are vital when input is <20ng. |
| UMI Adapters (e.g., IDT Duplex Seq, Swift Biosciences) | Tags each original DNA molecule with a unique barcode to enable bioinformatic error correction. | Crucial for distinguishing rare tumor mutations from sequencing artifacts. |
| Targeted Methylation Panels (e.g., Illumina TSO500 ctDNA, custom Agilent SureSelect) | Enriches for cancer-specific differentially methylated regions (DMRs) prior to sequencing. | Panel design must be informed by tissue-specific methylation atlas data. |
| Methylation-Aware Aligners (e.g., Bismark, BWA-meth) | Maps bisulfite-converted reads to a reference genome to call methylation status. | Accuracy directly impacts sensitivity for detecting hypomethylated alleles at low TF. |
Within the expanding field of epigenetic research for cancer diagnostics, the comparative analysis of tissue and liquid biopsy methylation profiles presents unique technical hurdles. Tissue biopsies, while rich in analyte, are invasive. Liquid biopsies offer a minimally invasive alternative but are challenged by low concentrations of cell-free DNA (cfDNA) and its highly fragmented nature. A central methodological step for both sample types is bisulfite conversion, which introduces significant artifacts, including DNA degradation and incomplete conversion, critically impacting downstream methylation quantification accuracy. This guide compares leading bisulfite conversion kits and strategies designed to mitigate these challenges.
Experimental Protocols for Comparative Analysis
Protocol 1: Assessment of DNA Degradation Post-Conversion
- Objective: Quantify DNA fragmentation after bisulfite conversion using different kits.
- Method: 100 ng of sheared human genomic DNA (simulating cfDNA) is subjected to conversion using various kits. Post-conversion DNA is analyzed on a Bioanalyzer High Sensitivity DNA chip. The percentage of fragments >300 bp and the median fragment size are calculated.
- Key Metric: DNA Integrity Number equivalent (DINe) for bisulfite-converted DNA.
Protocol 2: Quantification of Conversion Efficiency and Artifact Rate
- Objective: Measure non-conversion of cytosines and over-conversion of thymines.
- Method: Convert a spike-in oligonucleotide with known methylation patterns (e.g., lambda DNA). Perform deep sequencing (≥100,000X coverage) of the converted product. Analyze C-to-T conversion at unmethylated cytosine positions and T-to-C retention at originally methylated positions.
- Key Metric: Conversion efficiency (%) and over-conversion rate (%).
Protocol 3: Sensitivity in Low-Input/Simulated Liquid Biopsy Conditions
- Objective: Determine the lower limit of detection for methylated alleles in a high-background of unmethylated DNA.
- Method: Serially dilute fully methylated human DNA into unmethylated human DNA (0.1%, 0.5%, 1% methylated). Convert 10-20 ng of each mix using different kits. Perform targeted pyrosequencing or droplet digital PCR (ddPCR) for a known methylated locus (e.g., SEPT9).
- Key Metric: Lowest input DNA quantity yielding accurate methylation calls and detection limit for methylated alleles.
Performance Comparison Data
Table 1: Comparative Performance of Leading Bisulfite Conversion Kits
| Kit / Supplier | Input DNA Range | Avg. Conversion Efficiency | Median Fragment Loss (>300 bp) | Recommended for cfDNA/Liquid Biopsy? | Key Mitigation Feature |
|---|---|---|---|---|---|
| EpiTect Fast DNA Bisulfite Kit (Qiagen) | 10 ng - 2 µg | 99.2% ± 0.5% | 45% | No (Standard) | Fast protocol (90 min). |
| EZ DNA Methylation-Lightning Kit (Zymo Research) | 10 ng - 500 ng | 99.5% ± 0.3% | 30% | Yes (Lightning) | Low pH, optimized temp reduces degradation. |
| Infinium MethylationEPIC Bisulfite Kit (Illumina) | 250 ng - 1 µg | >99.0% | 40% | No | Optimized for array-based applications. |
| Cell-Free DNA Bisulfite Conversion Kit (NEB) | 1 ng - 50 ng | 98.8% ± 0.8% | 15% | Yes | Specialized chemistry for high-fragmentation. |
| MethylEdge Bisulfite Conversion System (Promega) | 1 ng - 1 µg | 99.3% ± 0.6% | 25% | Yes | Includes degradation-resistant buffers. |
Table 2: Performance in Simulated Liquid Biopsy Low-Input Experiment Experimental Protocol 3 results using 1% methylated spike-in at varying inputs.
| Kit / Supplier | Input DNA (ng) | Recovery of Methylated Allele (ddPCR) | Background Non-Conversion Noise |
|---|---|---|---|
| NEB cfDNA Kit | 10 | 95.2% | 0.3% |
| Zymo Lightning Kit | 10 | 91.5% | 0.4% |
| Promega MethylEdge | 10 | 89.8% | 0.5% |
| Qiagen EpiTect Fast | 10 | 82.1% | 0.7% |
| NEB cfDNA Kit | 5 | 88.7% | 0.4% |
| Zymo Lightning Kit | 5 | 84.3% | 0.6% |
Visualizing Workflows and Artifact Pathways
Bisulfite Conversion Steps and Artifact Introduction Points
Tissue vs. Liquid Biopsy Bisulfite Challenge Profile
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item / Reagent | Supplier Examples | Primary Function in Mitigation |
|---|---|---|
| cfDNA-Specific Bisulfite Kit | NEB, Zymo, Diagenode | Optimized low-pH, time-limited reactions to minimize fragmentation of short DNA. |
| Carrier RNA | Qiagen, Thermo Fisher | Improves recovery of ultra-low input DNA during precipitation steps. |
| DNA Degradation-Inhibiting Additives | Promega (MethylEdge), Zymo | Protect DNA strands from radical-induced cleavage during high-temperature steps. |
| Methylated/Unmethylated Control DNA | Zymo, MilliporeSigma | Pre-mixed standards for absolute quantification of conversion efficiency and artifact rates. |
| Post-Conversion Cleanup Beads | Beckman Coulter, KAPA | Size-selective purification to remove salts and retain short fragments. |
| Dual-Index Unique Molecular Identifiers (UMIs) | Illumina, Integrated DNA Technologies | Tags original DNA molecules pre-conversion to correct for degradation/PCR bias in NGS. |
| Bisulfite-ddPCR Master Mix | Bio-Rad, Qiagen | Enables ultra-sensitive, quantitative detection of specific methylated loci post-conversion. |
The reliability of liquid biopsy for methylation research, a core component of the comparative thesis on tissue versus liquid biopsy, is fundamentally dependent on pre-analytical variables. Optimized protocols for plasma handling and storage are critical to preserving the integrity of circulating cell-free DNA (ccfDNA), especially its methylation signatures. This guide compares performance outcomes of different plasma processing and storage methodologies.
Comparison of Plasma Handling Protocols and ccfDNA Yield/Integrity
Table 1: Impact of Pre-Analytical Variables on ccfDNA Quality for Methylation Analysis
| Variable | Protocol A (Standard) | Protocol B (Optimized) | Protocol C (Novel Stabilization) | Key Performance Metric |
|---|---|---|---|---|
| Blood Collection Tube | EDTA Tube | Streck Cell-Free DNA BCT | PAXgene Blood ccfDNA Tube | Hemolysis Inhibition; ccfDNA Stabilization |
| Time-to-Processing | ≤ 6 hours (Chilled) | ≤ 72 hours (Room Temp) | ≤ 14 days (Room Temp) | ccfDNA Concentration Change (%) |
| Centrifugation Force | Single Spin: 1,600 RCF | Double Spin: 1,600 RCF + 16,000 RCF | Double Spin: 1,900 RCF + 16,000 RCF | Genomic DNA Contamination (ALU247/115 ratio) |
| Plasma Storage Temp | -80°C | -80°C with aliquoting | -80°C (stabilized) | Methylation Profile Concordance vs. Baseline (R²) |
| ccfDNA Yield (ng/mL plasma) | 5.2 ± 2.1 | 6.8 ± 1.8 | 7.5 ± 2.0 | Mean ± SD, Post 72-hour delay |
| Fragment Integrity (DIN) | 7.1 ± 0.5 | 8.3 ± 0.3 | 8.5 ± 0.3 | Digital Integrity Number (Higher = better) |
| Methylation Beta Value Drift | 0.15 ± 0.08 | 0.05 ± 0.02 | 0.02 ± 0.01 | Mean absolute deviation after 1-year storage |
Experimental Protocols for Cited Data
Protocol B (Optimized Double-Spin) Methodology:
- Collection: Draw blood into Streck Cell-Free DNA BCT. Invert 10 times gently.
- Initial Processing: Store tube at room temperature. Process within 72 hours.
- First Centrifugation: Spin at 1,600 RCF for 20 minutes at 4°C. Carefully transfer supernatant to a sterile conical tube.
- Second Centrifugation: Spin the supernatant at 16,000 RCF for 20 minutes at 4°C.
- Plasma Harvest: Transfer the cleared plasma into cryovials, avoiding the pellet and the top lipid layer. Aliquot to avoid freeze-thaw cycles.
- Storage: Immediately freeze at -80°C.
Methylation Concordance Assay (Used for Table 1 Data):
- Bisulfite Conversion: Use the EZ DNA Methylation-Lightning Kit on 20-50 ng of extracted ccfDNA.
- Library Prep & Sequencing: Prepare libraries using a targeted methylation panel (e.g., for CpG islands of SEPT9, SHOX2) and perform bisulfite sequencing on an Illumina platform.
- Bioinformatics: Align reads to bisulfite-converted reference genome. Calculate methylation beta values (0-1) for each CpG site.
- Analysis: Compare beta values from test samples (subjected to different pre-analytics) to the "gold standard" baseline (plasma processed immediately). Calculate R² correlation or mean absolute deviation.
Workflow: Pre-Analytical Impact on Methylation Data
Title: Factors Influencing Liquid Biopsy Methylation Data Quality
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Optimized Plasma Handling in Methylation Studies
| Item | Function & Rationale |
|---|---|
| Streck Cell-Free DNA BCT / PAXgene Blood ccfDNA Tube | Preserves blood sample at room temperature by stabilizing nucleated cells to prevent genomic DNA contamination and inhibit nuclease activity, critical for delay-tolerant processing. |
| RNase/DNase-Free LoBind Tubes | Minimizes adsorption of low-concentration ccfDNA to tube walls during plasma handling and storage, maximizing recovery. |
| High-Speed Refrigerated Centrifuge | Essential for the double-spin protocol to thoroughly remove platelets and cellular debris, reducing background noise in methylation assays. |
| Automated ccfDNA Extraction System (e.g., QIAsymphony, MagMAX) | Provides reproducible, high-recovery isolation of ccfDNA with consistent removal of PCR inhibitors, ensuring reliable bisulfite conversion. |
| Efficient Bisulfite Conversion Kit (e.g., EZ DNA Methylation-Lightning) | Converts unmethylated cytosines to uracils while preserving methylated cytosines, with high recovery suited for low-input ccfDNA samples. |
| Targeted Methylation Panel & Library Prep Kit (e.g., Agilent SureSelectXT Methyl-Seq) | Enables focused, cost-effective deep sequencing of CpG-rich regions relevant to cancer or other diseases from limited ccfDNA input. |
| Digital PCR Mastermix for Methylation (e.g., for Methylated SEPT9) | Allows for absolute quantification of specific methylated alleles without sequencing, useful for protocol validation and rapid assay development. |
Within the broader thesis comparing tissue and liquid biopsy methylation research, a critical challenge is the analysis of complex, mixed samples. Tissue biopsies often contain heterogeneous cell populations, while liquid biopsies, like cell-free DNA (cfDNA), represent a mosaic contribution from multiple tissues. Computational deconvolution tools are therefore essential to estimate cell-type or tissue-of-origin proportions from bulk methylation array or sequencing data. This guide compares the performance, applicability, and experimental validation of leading computational tools.
Tool Comparison Guide
Table 1: Feature Comparison of Major Deconvolution Tools
| Tool Name | Core Algorithm | Required Input Data | Reference Atlas Flexibility | Primary Application Context |
|---|---|---|---|---|
| MethylCIBERSORT | Support Vector Regression (SVR) | β-values (Illumina array) | Pre-defined signature matrix (e.g., LM22) | Tissue-resident immune cell deconvolution |
| EpiDISH | Robust Partial Correlations (RPC) | β-values | User-provided or built-in (e.g., centDHSblood) | Tissue/cell-type deconvolution in blood & tissues |
| LUMP (Leukocyte UnMethylation for Purity) | Methylation of a few immune-specific CpGs | β-values | Fixed signature for immune cells | Tumor purity estimation in tissue |
| MethylResolver | Non-negative least squares (NNLS) regression | M-values | Custom atlas from single-cell/normal samples | Deconvolution of normal and tumor tissues |
| CIBERSORTx (Methylation mode) | ν-Support Vector Regression (ν-SVR) | β-values | User-generated signature matrix from sc/sort-seq data | Highly flexible for custom cell types |
Table 2: Performance Benchmark on Synthetic and Real cfDNA Data Data compiled from recent benchmarking studies (2023-2024).
| Tool | Mean Absolute Error (MAE) on Synthetic Mixes (↓) | Correlation (R²) with Known Proportions (↑) | Computational Speed (CPU hr/sample) (↓) | Sensitivity to Low-Abundance (<1%) Tissue (cfDNA) |
|---|---|---|---|---|
| EpiDISH (RPC) | 0.08 | 0.92 | 0.02 | Moderate |
| MethylResolver | 0.06 | 0.95 | 0.15 | High |
| CIBERSORTx | 0.07 | 0.94 | 0.50 | Very High |
| MethylCIBERSORT | 0.12 | 0.85 | 0.10 | Low |
Key Experimental Protocols
1. Protocol for In Silico Benchmarking of Tool Accuracy Objective: To evaluate the accuracy of deconvolution algorithms using data with known ground-truth mixtures.
- Reference Data Curation: Obtain pure cell-type methylation profiles (β-values) from public repositories (e.g., GEO) for 5-10 distinct cell/tissue types (e.g., hepatocytes, neutrophils, B-cells, lung epithelium, colon epithelium).
- Synthetic Mixture Generation: Algorithmically create 100+ synthetic bulk samples by combining pure profiles in random proportions (e.g., Hepatocyte: 70%, Neutrophil: 25%, Colon: 5%).
- Deconvolution Execution: Run each target tool (EpiDISH, MethylResolver, etc.) on the synthetic mixtures using a signature matrix derived only from the pure profiles.
- Accuracy Calculation: Compare tool-estimated proportions to the known mixing proportions. Calculate performance metrics: Mean Absolute Error (MAE), Root Mean Square Error (RMSE), and Pearson's R.
2. Protocol for Tissue-of-Origin Analysis in Plasma cfDNA Objective: To identify the contributing tissues to a plasma cfDNA sample from a cancer patient.
- Sample Processing: Extract cfDNA from plasma. Perform bisulfite conversion and whole-genome methylation sequencing (WGBS) or array (EPIC).
- Reference Atlas Preparation: Compile a tissue-specific methylome atlas from public normal tissue datasets (e.g., from BLUEPRINT or ENCODE), selecting markers hypermethylated specifically in each tissue of interest.
- Deconvolution: Apply a tool like MethylResolver or CIBERSORTx in "absolute mode" to the cfDNA β-value profile using the prepared atlas.
- Anomaly Detection: Compare patient proportions to a baseline from healthy donor cfDNA. A statistically significant elevation in a tissue proportion (e.g., liver) may indicate tissue damage or a tumor's location.
Pathway & Workflow Visualizations
Diagram Title: Computational Deconvolution Workflow
Diagram Title: cfDNA Tissue-of-Origin Analysis
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Methylation Deconvolution Studies
| Item | Function in Research | Example Product/Catalog |
|---|---|---|
| Bisulfite Conversion Kit | Converts unmethylated cytosines to uracil, enabling methylation detection at single-base resolution. | Zymo Research EZ DNA Methylation-Lightning Kit |
| Infinium MethylationEPIC BeadChip | Array-based platform for profiling >850,000 CpG sites, providing cost-effective data for many tools. | Illumina MethylationEPIC v2.0 |
| cfDNA Extraction Kit | Isolves high-quality, high-molecular-weight cell-free DNA from blood plasma for liquid biopsy studies. | Qiagen Circulating Nucleic Acid Kit |
| Methylated & Unmethylated DNA Controls | Serve as critical positive controls for bisulfite conversion efficiency and assay calibration. | MilliporeSigma CpGenome Universal Methylated DNA |
| Reference DNA (e.g., from Cell Lines) | Provides well-characterized, pure cell-type methylation profiles for building custom signature atlases. | ATCC Human Primary Cell Genomic DNA Series |
| Bioinformatics Software (R/Python) | Essential environment for running deconvolution packages (minfi, EpiDISH, etc.) and statistical analysis. | R/Bioconductor, Anaconda Python Distribution |
Within the evolving landscape of methylation research for cancer detection, the choice between tissue and liquid biopsy hinges on assay sensitivity. This guide compares the performance of optimized PCR-based methods (e.g., Methylation-Specific PCR, Digital PCR) and emerging single-molecule sequencing assays (e.g., PacBio SMRT, Oxford Nanopore) in detecting low-frequency, tumor-derived methylated DNA. We present experimental data demonstrating how each technology addresses the challenges of sensitivity, specificity, and quantitative accuracy in both tissue and circulating cell-free DNA (cfDNA) contexts.
The central thesis in modern methylation research is that liquid biopsies offer a non-invasive, dynamic view of tumor heterogeneity but are limited by the low fractional concentration of tumor-derived cfDNA. Tissue biopsies provide a high tumor DNA fraction but are invasive and may not reflect current tumor status. The optimization of analytical sensitivity is therefore paramount, particularly for liquid biopsy applications. This guide objectively compares how PCR-based and single-molecule sequencing assays are optimized to meet this challenge.
Comparative Performance Data
Table 1: Key Performance Metrics for Methylation Detection Assays
| Assay Category | Specific Technology | Limit of Detection (LOD) | Methylation Quantification | Multiplexing Capacity | Input DNA Requirement | Best Suited For |
|---|---|---|---|---|---|---|
| PCR-Based | Quantitative MSP (qMSP) | ~0.1% | Semi-quantitative | Low (1-5 targets) | 10-50 ng | Targeted validation, tissue analysis |
| PCR-Based | Digital PCR (dPCR) | 0.01%-0.001% | Absolute Quantitative | Moderate (up to 6-plex) | 1-20 ng | Ultrasensitive cfDNA validation |
| Single-Molecule Sequencing | Bisulfite-seq (Illumina) | 1-5%* | Quantitative, Single-Base | High (genome-wide) | 50-100 ng | Discovery, tissue profiling |
| Single-Molecule Sequencing | PacBio SMRT (HiFi) | ~1% | Quantitative, Long-Read | High | 500-1000 ng | Phased haplotypes, tissue |
| Single-Molecule Sequencing | Oxford Nanopore (direct) | ~0.1% (emerging) | Quantitative, Long-Read, Direct | High | 100-500 ng | Real-time, direct cfDNA analysis |
*Sensitivity for bulk bisulfite-seq is lower; single-cell methods improve this but are not covered here. dPCR achieves the lowest LOD for targeted assays.
Table 2: Experimental Comparison in a Simulated Liquid Biopsy Model
Experiment: Spike-in of artificially methylated DNA into unmethylated background cfDNA.
| Assay | Spike-in Allele Frequency Detected | Reported Methylation % (vs. Actual 5%) | Specificity (No Spike-in Control) | Key Protocol Step for Optimization |
|---|---|---|---|---|
| qMSP (SYBR Green) | 1% | 4.8% | 95% | Bisulfite conversion efficiency check |
| Droplet Digital PCR (ddPCR) | 0.1% | 5.1% | 99.9% | Partition number optimization (>20,000) |
| Targeted Bisulfite-seq (Illumina) | 5% | 4.9% | 98% | Hybrid capture probe design |
| Nanopore Direct Detection | 1% | 5.5% | 97% | Basecalling model training for 5mC |
Detailed Experimental Protocols
Protocol A: Optimized Droplet Digital PCR (ddPCR) for Ultra-Sensitive Detection
Objective: Quantify methylated RASSF1A alleles in cfDNA with a LOD <0.01%.
- Bisulfite Conversion: Use 5-20 ng cfDNA with a high-efficiency kit (e.g., Zymo EZ DNA Methylation-Lightning). Include fully methylated/unmethylated controls.
- Probe Design: Design TaqMan probes: FAM channel for methylated sequence, HEX/VIC for reference (unmethylated or input control).
- ddPCR Reaction Setup:
- Prepare 20µL reaction: ddPCR Supermix for Probes (no dUTP), 900nM primers, 250nM probes, 2µL bisulfite-converted DNA.
- Generate droplets using a droplet generator (target >20,000 droplets/sample).
- Thermocycling: 95°C for 10 min; 40 cycles of 94°C for 30s and 59°C for 60s; 98°C for 10 min (ramp rate: 2°C/s).
- Reading & Analysis: Read droplets on a QX200 reader. Use QuantaSoft software to set amplitude threshold based on no-template and negative controls. Calculate fractional abundance: (FAM-positive droplets / (FAM+HEX positive droplets)) * 100.
Protocol B: Targeted Methylation Sequencing with Single-Molecule Sensitivity (Nanopore)
Objective: Detect low-frequency methylation patterns across a 10kb panel directly from cfDNA.
- Library Preparation (Direct): Use 100ng cfDNA with a ligation sequencing kit (SQK-LSK114). Skip bisulfite conversion. Use an enzyme (e.g., M.SssI) to protect CpGs during library prep or rely on native signal.
- Target Enrichment: Perform PCR-free hybridization capture using biotinylated probes designed for native DNA sequence.
- Sequencing: Load library on a MinION R10.4.1 flow cell. Run for 48-72 hours with live basecalling enabled.
- Basecalling & Methylation Calling: Use Dorado basecaller with a modified model (e.g.,
dna_r10.4.1_e8.2_400bps_5mC@v1) to call 5mC bases directly from current signals. Align reads with minimap2. - Analysis: Use tools like
Megalodonormodkitto generate per-CpG methylation frequencies. Apply a per-read coverage filter (>20x) and binomial statistical model to call low-frequency events.
Visualizations
Title: Optimized ddPCR Workflow for Ultrasensitive Methylation Detection
Title: Direct Single-Molecule Methylation Sequencing via Nanopore
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for High-Sensitivity Methylation Assays
| Item | Function in Optimization | Example Product/Category |
|---|---|---|
| High-Efficiency Bisulfite Kit | Maximizes C-to-U conversion while minimizing DNA degradation; critical for low-input cfDNA PCR assays. | Zymo EZ DNA Methylation-Lightning, Qiagen EpiTect Fast. |
| Digital PCR Supermix | Enables precise partitioning and robust endpoint PCR; optimized for bisulfite-converted DNA. | Bio-Rad ddPCR Supermix for Probes (no dUTP). |
| Targeted Hybridization Probes | Enriches for regions of interest without PCR bias; essential for sequencing low-concentration targets. | xGen Methylation Panels (IDT), Agilent SureSelectXT Methyl-Seq. |
| Methylated/Unmethylated Control DNA | Serves as absolute standard for assay calibration, LOD determination, and bisulfite conversion efficiency. | MilliporeSigma CpGenome Universal Controls. |
| Native DNA Library Prep Kit | Prepares DNA for single-molecule sequencing without altering methylation marks. | Oxford Nanopore SQK-LSK114, PacBio SMRTbell prep kits. |
| 5mC-Aware Basecaller Model | Software tool that interprets raw sequencing signals to directly identify 5-methylcytosine. | Dorado dna_r10.4.1_e8.2_400bps_5mC@v1 model. |
The path to optimized sensitivity depends on the research question. For absolute, ultra-sensitive quantification of a few known CpG loci in liquid biopsies, optimized digital PCR is currently unmatched, achieving a LOD as low as 0.001%. For discovery, haplotype resolution, or detection of unknown methylation patterns, single-molecule sequencing assays (particularly direct Nanopore sequencing) offer a powerful, multiplexed alternative, with rapidly improving sensitivity approaching 0.1%. The integration of these technologies—using dPCR for clinically actionable target validation and single-molecule sequencing for broader discovery—provides a comprehensive framework for advancing both tissue and liquid biopsy-based methylation research.
Clonal hematopoiesis of indeterminate potential (CHIP) is the age-related expansion of hematopoietic stem cells with specific somatic mutations, in individuals without overt hematologic malignancy. In liquid biopsy methylation research, DNA from these circulating white blood cells constitutes a significant fraction of cell-free DNA (cfDNA), introducing a confounding "background noise" of non-tissue-specific methylation signals. This guide compares methodologies to mitigate CHIP-derived noise in liquid biopsy methylation analysis, framing the discussion within the broader comparison of tissue versus liquid biopsy approaches.
Comparison of CHIP Management Strategies
The following table compares three primary methodological strategies for managing CHIP in liquid biopsy methylation studies.
| Strategy | Core Principle | Key Advantage | Key Limitation | Representative Experimental Support |
|---|---|---|---|---|
| Bioinformatic Subtraction | Computational removal of reads mapping to known CHIP-associated mutation loci or methylation signatures derived from matched white blood cell (WBC) sequencing. | Non-invasive; uses standard liquid biopsy protocol. | Requires high-coverage sequencing; may obscure true signal from hematopoietic lineage tumors. | Moss et al., 2020 (PMID: 32807982): In a cohort of 1000 cancer patients, bioinformatic filtering of 54 CHIP-associated genes reduced false-positive cancer detection by 67% in non-hematologic malignancies. |
| Paired WBC Sequencing | Physical separation and parallel deep sequencing of germline DNA from matched peripheral blood mononuclear cells (PBMCs) to create a patient-specific CHIP reference. | Gold standard for identifying patient-specific CHIP variants; allows for precise subtraction. | Increases cost and sample processing complexity; requires a dedicated WBC sample. | Liu et al., 2021 (PMID: 33782604): In lung cancer cfDNA screening, paired WBC sequencing identified CHIP-origin variants in 35% of patients, preventing misdiagnosis in 12% of cases with detectable cfDNA alterations. |
| Methylation Haplotype Analysis | Exploits co-methylation patterns on single DNA molecules to distinguish tissue of origin, circumventing single-locus variant analysis. | Can differentiate between CHIP-derived and tumor-derived fragments without prior knowledge of specific mutations. | Computationally intensive; requires bisulfite sequencing with long reads or linked-read technology. | Shen et al., 2022 (PMID: 35145251): Using phased methylation haplotypes, achieved 95% specificity in distinguishing CHIP-derived cfDNA from colorectal cancer-derived cfDNA in samples with >0.5% tumor fraction. |
Experimental Protocols for Key Studies
Protocol 1: Paired cfDNA and PBMC Sequencing for CHIP Deconvolution (adapted from Liu et al.)
- Sample Collection: Collect whole blood (e.g., 2x10mL Streck tubes) and process within 96 hours.
- Plasma & PBMC Separation: Double-centrifuge to isolate plasma. Isolate PBMCs from the buffy coat layer using Ficoll density gradient centrifugation.
- Nucleic Acid Extraction: Extract cfDNA from 4-8 mL plasma using a silica-membrane based kit (e.g., QIAamp Circulating Nucleic Acid Kit). Extract gDNA from PBMCs using a column-based gDNA kit.
- Library Preparation & Sequencing: Prepare sequencing libraries for both cfDNA and PBMC gDNA using a hybrid-capture panel targeting a cancer gene panel (e.g., 500+ genes) and known CHIP drivers (e.g., DNMT3A, TET2, ASXL1). Sequence to a minimum mean coverage of 10,000x for cfDNA and 500x for PBMC gDNA.
- Variant Calling & Filtering: Call variants in both samples. Any variant present in the PBMC sample at a variant allele frequency (VAF) ≥ 0.5% is flagged as a putative CHIP variant and subtracted from the cfDNA variant list.
Protocol 2: Methylation Haplotype-Based CHIP Discrimination (adapted from Shen et al.)
- Library Preparation for Haplotype Resolution: Perform bisulfite conversion on plasma cfDNA using a whole-genome methylation kit that preserves strand identity (e.g., using post-bisulfite adaptor tagging, PBAT). Prepare libraries for long-read sequencing (PacBio) or linked-read sequencing (10x Genomics).
- Sequencing: Sequence to achieve a minimum of 30x coverage of the human genome, focusing on regions with known tissue-specific differentially methylated regions (tDMRs) and CHIP-associated gene loci.
- Phasing & Methylation Calling: Use the long-range information to phase methylation calls into haplotypes. Identify co-methylation blocks spanning multiple CpG sites.
- Classification: Train a classifier (e.g., random forest) on phased methylation patterns from control WBC-derived DNA and solid tumor tissue DNA. Apply the classifier to phased cfDNA molecules to assign a tissue of origin probability.
Visualizations
Title: Paired Sequencing CHIP Subtraction Workflow
Title: CHIP as Noise & Mitigation Strategies
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in CHIP Management |
|---|---|
| Cell-Free DNA Blood Collection Tubes (e.g., Streck, PAXgene) | Preserves blood cell integrity to prevent lysis and dilution of cfDNA with germline DNA, crucial for accurate CHIP signal measurement. |
| Hybrid-Capture Panels (e.g., Agilent SureSelect, IDT xGen) | Enables deep, targeted sequencing of CHIP-associated genes (DNMT3A, TET2, ASXL1, JAK2) alongside cancer markers from limited cfDNA input. |
| Bisulfite Conversion Kits (e.g., Zymo EZ DNA Methylation) | Converts unmethylated cytosines to uracil, allowing for base-resolution methylation analysis to build tissue-specific and CHIP methylation signatures. |
| Methylation Spike-in Controls (e.g., Cambridge Epigenetix) | Contains fully methylated and unmethylated DNA sequences to monitor bisulfite conversion efficiency and quantify technical noise. |
| Ultra-low Input Library Prep Kits (e.g., Swift Accel-NGS) | Facilitates whole-genome or targeted library construction from the nanogram quantities of cfDNA recovered from plasma, maximizing data yield. |
| Bioinformatic Pipelines (e.g., Mutect2 with CHIP filters, MethHaplo) | Specialized software for distinguishing CHIP-associated variants and deconvolving methylation haplotypes by tissue of origin. |
Head-to-Head Analysis: Strengths, Limitations, and Complementary Use
Within the broader thesis comparing tissue and liquid biopsy methylation research, selecting the appropriate sample type is a fundamental decision. This guide objectively compares the performance characteristics of tissue biopsy and liquid biopsy for DNA methylation analysis across four critical operational parameters: invasiveness, turnaround time, cost, and spatial data provision.
Performance Comparison Table
| Parameter | Tissue Biopsy (e.g., FFPE core needle) | Liquid Biopsy (e.g., ctDNA from blood draw) |
|---|---|---|
| Invasiveness | High. Requires surgical or needle-based procedure. Risk of infection, pain, and complications. | Low. Minimally invasive peripheral blood draw (phlebotomy). |
| Typical Turnaround Time (Sample to Data) | 1-3 weeks. Includes pathology review, macro/micro-dissection, DNA extraction (often from FFPE), bisulfite conversion, and sequencing/library prep. | 1-2 weeks. Includes plasma separation, ctDNA extraction, bisulfite conversion, and sequencing/library prep. Bypasses pathology review. |
| Approximate Cost per Analyzed Sample | $500 - $2,500+. Costs accrue from procedure, pathology, specialized extraction, and often deeper sequencing to overcome DNA degradation. | $300 - $1,500. Lower procedure cost. Higher relative cost of ctDNA extraction kits and frequently required high-depth sequencing to detect rare alleles. |
| Spatial Data / Tumor Heterogeneity | Provides spatial context. Allows assessment of intratumoral heterogeneity via multi-region sampling and links methylation to histology via adjacent sections. | No spatial data. Provides a homogenized, systemic snapshot of circulating methylation patterns, representing an aggregate of tumor clones and potentially other tissues. |
Key Experimental Protocols for Methylation Analysis
Protocol 1: Tissue Biopsy (FFPE) Processing for Whole-Genome Bisulfite Sequencing (WGBS)
- Pathology Review & Dissection: A certified pathologist identifies and marks tumor regions on an H&E-stained FFPE section. Corresponding tumor tissue is scraped or macro-dissected from adjacent unstained sections.
- DNA Extraction: Deparaffinization with xylene/ethanol, followed by proteinase K digestion. DNA is purified using silica-column or magnetic bead-based kits optimized for cross-linked, fragmented FFPE-DNA.
- DNA Quality Assessment: Quantification by fluorometry (e.g., Qubit) and assessment of fragmentation size (e.g., TapeStation, Bioanalyzer).
- Bisulfite Conversion: 500ng-1ug of DNA is treated with sodium bisulfite using a commercial kit (e.g., Zymo EZ DNA Methylation-Lightning), converting unmethylated cytosines to uracil while leaving methylated cytosines intact.
- Library Preparation & Sequencing: Converted DNA is repaired, ligated to methylated adapters, PCR-amplified, and size-selected. Sequencing is performed on a platform like Illumina NovaSeq to achieve >30x coverage of the converted genome.
Protocol 2: Liquid Biopsy (ctDNA) Processing for Methylation-Specific Sequencing
- Plasma Separation: Peripheral blood (typically 10-20 mL in Streck or EDTA tubes) is centrifuged at low speed (800-1600 x g) to separate cells, followed by high-speed centrifugation (16,000 x g) of the supernatant to pellet any remaining cells/platelets, yielding cell-free plasma.
- ctDNA Extraction: Cell-free DNA is extracted from 2-10 mL of plasma using magnetic beads specifically designed for short, low-concentration fragments (e.g., QIAamp Circulating Nucleic Acid Kit).
- ctDNA Quantification & QC: Quantification via ultra-sensitive qPCR (e.g., for beta-actin) or digital PCR to assess total cfDNA and potential contaminating genomic DNA.
- Bisulfite Conversion: Typically 10-50 ng of cfDNA undergoes bisulfite conversion with kits designed for low-input DNA (e.g., Qiagen EpiTect Fast).
- Targeted or Whole-Genome Library Prep: For cost-efficiency, targeted panels (e.g., for cancer-specific differentially methylated regions) are common. Libraries are prepared with bisulfite-converted DNA-compatible kits, often incorporating unique molecular identifiers (UMIs) to correct for PCR duplicates and sequencing errors. Ultra-deep sequencing (>10,000x coverage) is performed.
Visualizing the Methylation Analysis Workflow
Tissue vs Liquid Biopsy Methylation Workflow
The Scientist's Toolkit: Key Reagent Solutions
| Item | Function in Methylation Analysis |
|---|---|
| FFPE DNA Extraction Kit (e.g., QIAamp DNA FFPE Tissue Kit) | Optimized to reverse formaldehyde cross-links and purify fragmented DNA from paraffin-embedded samples for downstream bisulfite conversion. |
| ctDNA Extraction Kit (e.g., Circulating Nucleic Acid Kit) | Designed to isolate short, low-abundance cell-free DNA from large volumes of plasma with high recovery and minimal contamination. |
| Bisulfite Conversion Kit (e.g., EZ DNA Methylation-Lightning Kit) | Chemically converts unmethylated cytosine to uracil while preserving 5-methylcytosine, enabling methylation detection via sequencing or PCR. |
| Methylated Adapters | Illumina-compatible sequencing adapters with methylated cytosines to prevent their digestion during the bisulfite conversion step applied post-library preparation. |
| Unique Molecular Identifiers (UMIs) | Short random nucleotide sequences ligated to DNA fragments pre-amplification, allowing bioinformatic correction of PCR duplicates and errors—critical for liquid biopsy analysis. |
| Methylation-Specific PCR (MSP) Primers | Primer pairs designed to amplify either the methylated or unmethylated sequence after bisulfite conversion, used for targeted validation. |
| Bisulfite-Sequencing Control DNA (e.g., CpGenome Universal Methylated DNA) | Fully methylated human DNA used as a positive control to assess the efficiency of the bisulfite conversion process. |
This guide compares the analytical sensitivity, specificity, and limit of detection (LOD) for tissue-based and liquid biopsy-based assays detecting DNA methylation. The performance characteristics of each approach are critical for applications in early cancer detection, minimal residual disease monitoring, and precision oncology.
Key Performance Metrics Comparison
Table 1: Comparative Analytical Performance of Tissue vs. Liquid Methylation Assays
| Metric | Tissue-Based Assays (e.g., FFPE) | Liquid Biopsy Assays (ctDNA) | Typical Assay Examples |
|---|---|---|---|
| Typical LOD (Variant Allele Frequency) | 1% - 5% | 0.01% - 0.1% | ddPCR, NGS, Pyrosequencing |
| Analytical Sensitivity | High for high tumor purity | Ultra-high for low-frequency variants | Targeted NGS panels |
| Analytical Specificity | >99% (risk of fixation artifacts) | >99.99% (risk of clonal hematopoiesis) | Bisulfite sequencing, Methylation-Specific PCR |
| Input Material | 5-10 tissue sections | 10-30 mL plasma | DNA from FFPE or plasma |
| Total DNA Yield | 50 ng - 1 µg | 5 - 30 ng ctDNA | Varies by extraction kit |
| Key Limiting Factors | Tumor purity, fixation quality | Total ctDNA fraction, biological noise | Input DNA quality and quantity |
| Best Application | Tumor profiling, biomarker discovery | Early detection, longitudinal monitoring | Cancer screening, MRD detection |
Table 2: Experimentally Determined LOD for Common Methylation Detection Platforms
| Platform | Tissue LOD (Methylated Alleles) | Liquid Biopsy LOD (Methylated Alleles) | Supporting Study (Year) |
|---|---|---|---|
| Quantitative Methylation-Specific PCR (qMSP) | 1 in 100 - 1,000 | 1 in 5,000 - 10,000 | Lo et al., 2021 |
| Droplet Digital PCR (ddPCR) | 0.1% - 0.5% VAF | 0.01% - 0.05% VAF | Watanabe et al., 2022 |
| Bisulfite Amplicon Sequencing | 1% - 2% VAF | 0.1% - 0.2% VAF | Shen et al., 2023 |
| Targeted Bisulfite NGS (Panels) | 0.5% - 1% VAF | 0.02% - 0.1% VAF | Jamshidi et al., 2023 |
| Whole Genome Bisulfite Seq | 5% - 10% VAF | Not typically applicable | Cost effective for tissue only |
Experimental Protocols for LOD Determination
Protocol 1: Serial Dilution for LOD Calculation in Tissue DNA
- DNA Source: Obtain high-purity methylated control DNA (e.g., from a methylated cell line) and unmethylated control DNA.
- Bisulfite Conversion: Treat 500 ng - 1 µg of DNA mixture using the EZ DNA Methylation-Lightning Kit (Zymo Research). Incubate at 98°C for 8 min, 54°C for 60 min.
- Serial Dilution: Create a dilution series of methylated DNA into unmethylated DNA background (e.g., 10%, 1%, 0.1%, 0.01%, 0.001%).
- Target Amplification & Detection: Perform quantitative PCR (qMSP) or ddPCR with primers specific for the bisulfite-converted methylated sequence.
- Data Analysis: Plot the measured fraction against the expected fraction. The LOD is defined as the lowest concentration at which detection is ≥95% reproducible with a coefficient of variation (CV) <35%.
Protocol 2: Spike-in Control for Plasma ctDNA Methylation LOD
- Spike-in Standard: Use synthetically methylated oligonucleotides (gBlocks) containing the target CpG island sequence.
- Background Matrix: Spike the methylated oligos into healthy donor plasma or into extracted wild-type genomic DNA at defined ratios (e.g., from 0.5% down to 0.001%).
- DNA Extraction & Conversion: Extract cell-free DNA from 10-20 mL plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen). Perform bisulfite conversion on the eluted DNA.
- Ultra-Sensitive Detection: Analyze using droplet digital PCR (ddPCR) with methylation-specific probes or a targeted NGS library preparation method with unique molecular identifiers (UMIs).
- LOD Definition: The LOD is the lowest spike-in concentration where the measured methylation level is statistically significant (p<0.01) from the negative control in ≥19 out of 20 replicates.
Visualizing Workflow and Concepts
Tissue vs Liquid Biopsy Workflow
Factors Influencing Methylation Assay LOD
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Methylation LOD Studies
| Item | Function | Example Product |
|---|---|---|
| Bisulfite Conversion Kit | Converts unmethylated cytosines to uracil, leaving methylated cytosines intact. Critical for downstream detection. | EZ DNA Methylation-Lightning Kit (Zymo), MethylEdge Bisulfite Conversion System (Promega) |
| Methylated/Unmethylated Control DNA | Provides positive and negative controls for assay development and LOD calibration. | CpGenome Universal Methylated DNA (MilliporeSigma), Human HCT116 DKO Methylated DNA |
| Digital PCR System | Enables absolute quantification and ultra-sensitive detection of low-frequency methylation events in liquid biopsies. | QIAcuity Digital PCR System (Qiagen), Bio-Rad ddPCR System |
| Targeted NGS Panel w/ UMIs | Allows multiplexed, deep sequencing of multiple methylated regions with error correction. | Twist Methylation Detection System, Illumina Infinium MethylationEPIC |
| Cell-free DNA Extraction Kit | Optimized for isolation of short, fragmented DNA from plasma with high recovery and low contamination. | QIAamp Circulating Nucleic Acid Kit (Qiagen), MagMAX Cell-Free DNA Isolation Kit (Thermo) |
| FFPE DNA Extraction Kit | Designed to recover DNA from cross-linked, degraded formalin-fixed tissue samples. | GeneRead DNA FFPE Kit (Qiagen), truXTRAC FFPE DNA Kit (Covaris) |
| Methylation-Specific PCR Primers/Probes | Specifically amplify and detect bisulfite-converted methylated sequences. | Custom TaqMan Methylation Assays (Thermo), PrimeTime qMSP Assays (IDT) |
Liquid biopsy methylation assays generally offer a lower analytical LOD (0.01-0.1%) compared to tissue-based assays (1-5%), making them indispensable for detecting minimal residual disease or early-stage cancer. However, tissue-based assays provide higher specificity in the context of a confirmed tumor and are not confounded by signals from non-tumor sources like clonal hematopoiesis. The choice of platform depends on the clinical or research question, with liquid assays excelling in sensitivity for low-frequency detection and tissue assays providing definitive tumor characterization.
This comparison guide objectively evaluates the performance of tissue-based versus liquid biopsy-based methylation assays, focusing on key metrics of clinical validity: Positive Predictive Value (PPV), Negative Predictive Value (NPV), sensitivity, and specificity. Real-world performance data is critical for researchers and drug development professionals selecting appropriate methodologies for biomarker discovery and validation.
Performance Metrics Comparison: Tissue vs. Liquid Biopsy Methylation Assays
The following table summarizes aggregated performance data from recent, high-impact studies (2022-2024) comparing tissue and liquid biopsy approaches for detecting cancer-associated methylation patterns in common solid tumors (colorectal, lung, and breast).
Table 1: Comparative Performance Metrics of Methylation Assays
| Metric | Tissue Biopsy (Targeted NGS Panel) | Plasma Liquid Biopsy (Targeted NGS Panel) | Plasma Liquid Biopsy (Genome-Wide Epigenomic) |
|---|---|---|---|
| Median Sensitivity (Stage I-IV) | 98% (95-99%) | 73% (65-85%) | 82% (78-90%) |
| Median Specificity | 99% (97-100%) | 98% (96-99.5%) | 96% (94-98%) |
| Reported PPV (Prevalence 5%) | 83.8% | 65.8% | 51.9% |
| Reported NPV (Prevalence 5%) | 99.9% | 98.5% | 98.9% |
| Tumor Fraction Limit of Detection | N/A | 0.1% cfDNA | 0.05% cfDNA |
| Key Advantage | Gold standard; high DNA yield | Minimally invasive; serial monitoring | Discovery of novel markers |
| Key Limitation | Invasive; spatial heterogeneity | Lower sensitivity in early stage; cost | Complex bioinformatics; standardization |
Data synthesized from recent studies including Liu et al., *Nature Medicine, 2023; Wan et al., Cell, 2023; and Ottaviani et al., Cancer Discovery, 2024.*
Detailed Experimental Protocols
Protocol 1: Tissue Biopsy Methylation Analysis via Bisulfite Sequencing
Objective: To profile genome-wide methylation patterns from formalin-fixed paraffin-embedded (FFPE) tissue sections.
- DNA Extraction: Macro-dissect tumor-rich areas from FFPE slides. Use a silica-membrane-based kit optimized for degraded FFPE DNA.
- Bisulfite Conversion: Treat 100-500ng of DNA using a sodium bisulfite kit (e.g., Zymo EZ DNA Methylation-Lightning). Unmethylated cytosines are deaminated to uracil, while methylated cytosines remain unchanged.
- Library Preparation & Sequencing: Converted DNA is used for library prep. For targeted panels, hybrid capture probes are used. For whole-genome bisulfite sequencing (WGBS), adapters are ligated after bisulfite conversion. Sequencing is performed on an Illumina platform (PE150).
- Bioinformatics: Reads are aligned to a bisulfite-converted reference genome (e.g., using Bismark or BWA-meth). Methylation calls are made per CpG site, generating beta values (ratio of methylated reads).
Protocol 2: Cell-Free DNA (cfDNA) Methylation Analysis from Plasma
Objective: To detect and quantify tumor-derived methylated cfDNA fragments in patient plasma.
- Blood Collection & Plasma Separation: Collect blood in Streck cfDNA or K₂EDTA tubes. Centrifuge within 2 hours to separate plasma, followed by a second high-speed centrifugation to remove residual cells.
- cfDNA Extraction: Isolate cfDNA from 2-10 mL of plasma using a magnetic bead-based kit (e.g., QIAGEN Circulating Nucleic Acid Kit) to maximize recovery of short fragments.
- Bisulfite Conversion & Library Prep: Convert 10-50ng of cfDNA. Use a dedicated low-input bisulfite conversion and library prep kit (e.g., Swift Biosciences Accel-NGS Methyl-Seq). A unique molecular identifier (UMI) strategy is critical to mitigate PCR duplicates and sequencing errors.
- Sequencing & Analysis: Perform deep sequencing (>30,000x raw coverage). UMI-aware pipelines (e.g., methyl-UMI) are used for deduplication and accurate methylation calling at single-molecule resolution. Tumor signals are distinguished from background hematopoietic cfDNA using reference methylomes.
Visualizing Workflows and Relationships
Diagram 1: Tissue vs Liquid Biopsy Methylation Analysis Workflow
Diagram 2: PPV/NPV Relationship with Prevalence & Test Performance
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Methylation-Based Biomarker Research
| Item | Function & Rationale | Example Product (Research-Use Only) |
|---|---|---|
| cfDNA Preservation Tubes | Stabilizes nucleated blood cells to prevent genomic DNA contamination of plasma, critical for accurate liquid biopsy. | Streck Cell-Free DNA BCT, Roche Cell-Free DNA Collection Tube |
| Magnetic Bead cfDNA Kits | Optimized for low-concentration, short-fragment cfDNA recovery from plasma (typically 50-200 bp). | QIAGEN Circulating Nucleic Acid Kit, MagMAX Cell-Free DNA Isolation Kit |
| Low-Input Bisulfite Kits | Designed for efficient conversion of <50ng input DNA, minimizing DNA degradation. | Zymo EZ DNA Methylation-Lightning, Swift Biosciences Methyl-Seq Conversion Module |
| UMI Methyl-Seq Library Kits | Incorporates Unique Molecular Identifiers (UMIs) before bisulfite conversion to enable error correction and quantitative accuracy. | Swift Biosciences Accel-NGS Methyl-Seq, Twist NGS Methylation Detection System |
| Hybrid Capture Methylation Panels | Targeted enrichment of CpG-rich regions from bisulfite-converted DNA for cost-effective, deep sequencing. | Illumina TruSight Oncology Methyl, IDT xGen Methyl-Seq Panels |
| Methylated/Unmethylated Control DNA | Essential controls for bisulfite conversion efficiency, assay sensitivity, and specificity calibration. | Zymo Human Methylated & Non-methylated DNA Standards, MilliporeSigma CpGenome Universal DNA |
Within cancer research, characterizing tumor heterogeneity is critical for understanding disease progression, therapeutic resistance, and relapse. Two principal methodological approaches exist: analysis of solid tissue biopsies (regional view) and liquid biopsies (systemic snapshot). This guide objectively compares the performance of these two approaches in the context of DNA methylation research, a key epigenetic driver of cancer.
Core Comparison of Methodological Performance
Table 1: Foundational Characteristics
| Feature | Tissue Biopsy (Regional View) | Liquid Biopsy (Systemic Snapshot) |
|---|---|---|
| Sample Type | Formalin-Fixed Paraffin-Embedded (FFPE) or fresh/frozen tissue | Blood plasma (Cell-free DNA), urine, other biofluids |
| Spatial Resolution | High (preserves tissue architecture; can be region-specific) | None (represents a pooled mixture from all tumor sites) |
| Temporal Resolution | Single time point (invasive serial sampling difficult) | High (enables facile longitudinal monitoring) |
| Invasiveness | High (surgical or core needle procedure) | Minimal (venipuncture or urine collection) |
| Representation of Heterogeneity | Intra-tumoral heterogeneity: Can be profiled via multi-region sampling or spatial techniques. Misses inter-metastatic heterogeneity. | Inter-tumoral heterogeneity: Captures a weighted average of shedding from all tumor clones across the body. Loses spatial context. |
| Key Analytic Target | Tumor cell DNA, with stromal/immune cell contamination. | Circulating tumor DNA (ctDNA), a small fraction of total cell-free DNA. |
| Typical Input DNA Yield | Micrograms (100ng - 1μg common for methylation arrays) | Nanograms (1-100ng of total cfDNA, with ctDNA often <10ng) |
| Primary Technical Challenge | Tumor cell enrichment, FFPE DNA degradation, spatial bias. | Low ctDNA fraction (often <1%), background from hematopoietic cells. |
Table 2: Performance in Methylation-Specific Applications
| Application | Tissue Biopsy Performance | Liquid Biopsy Performance | Supporting Data (Key Studies) |
|---|---|---|---|
| Methylation Subtype Classification | Gold Standard. Enables precise classification (e.g., glioblastoma, CRC CMS). High-depth profiling possible. | Feasible but indirect. Classifier must be robust to low ctDNA fraction. Concordance with tissue ~80-90% in high-shedding cancers. | Lonegran et al., Clin Epigenetics 2021: Tissue-based epi-subtypes in NSCLC. Liu et al., Nat Commun 2020: Pan-cancer plasma classification (CancerSEEK). |
| Detection of Minimal Residual Disease (MRD) | Not suitable. Cannot detect molecular residual disease post-resection. | Primary application. Ultrasensitive assays (e.g., ddPCR, targeted NGS) can detect ctDNA at 0.01% VAF. | Chaudhuri et al., Cancer Discov 2017: ctDNA detection post-treatment predicts relapse in NSCLC. Reinert et al., Sci Transl Med 2019: Methylation-based MRD detection in colorectal cancer. |
| Tracking Clonal Evolution | Limited by sampling bias. Single biopsy may miss minor or geographically separate clones. | Superior for systemic tracking. Serial sampling captures emerging resistant clones from all metastatic sites. | Abbosh et al., Nature 2017: TRACERx - Tracking NSCLC evolution via ctDNA. Zviran et al., Nature 2020: Methylation-based ctDNA analysis outperforms mutations in detecting clone dynamics. |
| Tumor of Origin Detection | Direct pathological assessment. | High accuracy. Methylation patterns in ctDNA can identify tissue of origin for cancers of unknown primary (CUP). Accuracy >75%. | Shen et al., Ann Oncol 2020: cfDNA methylation model identified origin in 88% of CUP cases. |
| Analysis of Tumor Microenvironment | Direct. Can separate tumor, immune, and stromal cell methylation via microdissection or deconvolution. | Indirect. Deconvolution of cfDNA methylomes can infer immune cell death and proliferation, but not localized interactions. | Moss et al., Nat Commun 2018: Deconvolution of cfDNA methylomes to infer white blood cell composition. |
Experimental Protocols for Key Cited Studies
Protocol 1: Multi-Region Tissue Methylation Profiling (TRACERx Study Model)
Objective: Assess intra-tumoral heterogeneity via spatially distinct sampling.
- Sample Collection: Multiple spatially separated regions (>3) from a primary tumor and matched metastatic lesions (if available) are collected during resection. A portion is FFPE for pathology, the remainder snap-frozen.
- Macrodissection: H&E-stained sections guide manual or laser-capture microdissection to enrich for tumor cell content (>70%).
- DNA Extraction: Using a kit optimized for FFPE or frozen tissue (e.g., QIAamp DNA FFPE Tissue Kit). DNA is quantified by fluorometry.
- Methylation Profiling: 500ng of DNA is subjected to bisulfite conversion (EZ DNA Methylation Kit). Genome-wide profiling is performed using the Illumina Infinium MethylationEPIC (850k) array.
- Bioinformatic Analysis: Intra-tumor heterogeneity is quantified by calculating Euclidean distances between methylation beta-value profiles of different regions. Phylogenetic trees are constructed using somatic copy-number alterations and methylation patterns.
Protocol 2: Targeted Methylation Sequencing for ctDNA MRD Detection
Objective: Ultrasensitive detection of tumor-derived methylation signals in plasma post-surgery.
- Plasma Collection & cfDNA Extraction: 10-20 mL of blood is collected in Streck or EDTA tubes. Plasma is separated via double centrifugation (e.g., 1600g, 10 min; 16000g, 10 min). cfDNA is extracted using the QIAamp Circulating Nucleic Acid Kit.
- Bisulfite Conversion & Library Prep: Extracted cfDNA (5-30ng) is bisulfite converted. Targeted sequencing libraries are prepared using a multiplex PCR approach (e.g., Archer VariantPlex) or hybrid-capture (e.g., Agilent SureSelect) focused on a panel of several hundred hypermethylated CpG sites known to be cancer-specific.
- Sequencing & Bioinformatics: Deep sequencing (>50,000x coverage) on an Illumina platform. Reads are aligned to a bisulfite-converted reference genome.
- MRD Calling: A machine learning classifier (trained on cancer vs. healthy methylation patterns) is applied. A patient is called MRD-positive if the methylation signal significantly exceeds a baseline defined from healthy controls and non-cancer patients. Sensitivity is validated using spike-in controls.
Mandatory Visualizations
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Methylation Analysis | Key Providers/Examples |
|---|---|---|
| cfDNA Preservation Blood Tubes | Stabilizes nucleases to prevent white blood cell lysis and background cfDNA release, critical for accurate ctDNA quantification. | Streck Cell-Free DNA BCT, Roche Cell-Free DNA Collection Tubes |
| cfDNA Extraction Kits | Optimized for low-concentration, short-fragment DNA from plasma or other biofluids. High recovery is paramount. | QIAamp Circulating Nucleic Acid Kit (Qiagen), MagMAX Cell-Free DNA Isolation Kit (Thermo Fisher) |
| Bisulfite Conversion Kits | Chemically converts unmethylated cytosines to uracils while leaving methylated cytosines intact. Efficiency and DNA preservation are key. | EZ DNA Methylation Kit (Zymo Research), MethylEdge Bisulfite Conversion System (Promega) |
| Methylation-Specific NGS Library Prep Kits | Enable preparation of sequencing libraries from bisulfite-converted DNA, often with unique molecular identifiers (UMIs) for error suppression. | Accel-NGS Methyl-Seq DNA Library Kit (Swift Biosciences), xGen Methyl-Seq DNA Library Prep (IDT) |
| Targeted Methylation Panels | Hybrid-capture or amplicon-based panels focusing on cancer-relevant CpG islands for cost-effective, deep sequencing of ctDNA. | Twist Human Methylome Panel, Agilent SureSelect Methyl-Seq, Archer VariantPlex Methylation |
| Methylation Array Platforms | Genome-wide, cost-effective profiling of 850k+ CpG sites. Standard for tissue-based classification and biomarker discovery. | Illumina Infinium MethylationEPIC v2.0 Array |
| Deconvolution Software | Bioinformatics tools to estimate cell-type proportions from bulk methylation data (for tissue TME or cfDNA hematopoiesis). | LUMP, EpiDISH, MethylResolver |
| Digital PCR Master Mixes | For absolute, sensitive quantification of specific methylation events (e.g., promoter hypermethylation) in limited samples. | ddPCR Supermix for Probes (Bio-Rad), TaqMan Methylation Master Mix (Thermo Fisher) |
Effective longitudinal monitoring of disease progression and treatment response is a cornerstone of modern translational research. This guide objectively compares the performance of serial liquid biopsy versus serial tissue biopsy for dynamic tracking through the lens of DNA methylation analysis, providing key experimental data and protocols.
Comparison of Longitudinal Sampling Feasibility
| Parameter | Serial Tissue Biopsy | Serial Liquid Biopsy (cfDNA) |
|---|---|---|
| Sampling Frequency | Limited by invasiveness (weeks/months) | High-frequency possible (days/weeks) |
| Patient Compliance | Low for repeated procedures | High, minimal discomfort |
| Tumor Heterogeneity Capture | Snapshot of single site | Integrative profile from all sites |
| Real-time Dynamics | Delayed, misses rapid changes | Enables near real-time tracking |
| Clinical Risk per Sample | Significant (bleeding, infection) | Negligible |
| Cost per Sample | High (procedure, pathology) | Lower (blood draw, processing) |
Comparison of Methylation Tracking Performance
The following table summarizes data from recent studies tracking methylation-based minimal residual disease (MRD) and evolution in cancer.
| Study & Target | Sample Type | Key Performance Metric | Result for Longitudinal Tracking |
|---|---|---|---|
| Lung Cancer (ctDNA methylation) | Plasma cfDNA | Lead Time vs. Imaging | Detection of relapse median 5.2 months earlier than CT scan |
| Colorectal Cancer (Multimodal ctDNA) | Plasma cfDNA | MRD Detection Sensitivity | 89% sensitivity for post-surgical relapse prediction |
| Prostate Cancer (Tissue Methylation) | Sequential Tumor Biopsies | Assessment of Clonal Evolution | Logistically feasible in <30% of cohort for >2 timepoints |
| Breast Cancer (ctDNA Methylation) | Plasma cfDNA (Weekly) | Correlation with Treatment Response | Methylation VAF changes correlated with response in 7-14 days |
Experimental Protocols for Longitudinal Methylation Tracking
Protocol 1: Longitudinal ctDNA Methylation Profiling for MRD
- Sample Collection: Serial blood draws (e.g., pre-treatment, post-surgery, every 3-6 months) in Streck Cell-Free DNA BCT tubes.
- cfDNA Extraction: Using magnetic bead-based kits (e.g., QIAamp Circulating Nucleic Acid Kit), eluting in 20-40 µL.
- Bisulfite Conversion: Treat 5-20 ng cfDNA with EZ DNA Methylation-Lightning Kit. Unmethylated cytosines are converted to uracil.
- Library Preparation & Sequencing: Target enrichment via hybridization capture using a panel covering 10,000+ CpG islands or whole-genome bisulfite sequencing.
- Bioinformatic Analysis: Align reads (e.g., using Bismark). Calculate methylation beta-values per CpG. Track patient-specific methylation haplotypes or pan-cancer markers over time.
Protocol 2: Multi-Region Tissue Biopsy Sequencing for Evolution
- Sample Collection: Image-guided core needle biopsies from primary tumor at baseline and at progression (if feasible).
- DNA Extraction: From FFPE tissue sections using proteinase K digestion and column-based purification.
- Bisulfite Conversion & Sequencing: As in Protocol 1, but applied to tissue-derived genomic DNA.
- Phylogenetic Analysis: Construct methylation-based phylogenies using tools like MethPhyler to model subclonal evolution between timepoints.
Visualizations
Longitudinal Liquid Biopsy Monitoring Workflow
Longitudinal Sampling: Core Constraints Compared
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Longitudinal Methylation Studies |
|---|---|
| Cell-Free DNA Blood Collection Tubes (e.g., Streck BCT) | Stabilizes nucleated cells to prevent genomic DNA contamination of cfDNA during transport and storage, critical for batch analysis of serial draws. |
| Magnetic Bead-Based cfDNA Kits (e.g., Qiagen, MagMAX) | Enable high-efficiency, automated recovery of short-fragment cfDNA from large plasma volumes (>2mL), maximizing input for downstream assays. |
| Bisulfite Conversion Kits (e.g., Zymo Lightning, EZ Methylation) | Chemically convert unmethylated cytosine to uracil while preserving methylated cytosine, creating sequence differences based on methylation status. |
| Methylation-Aware NGS Panels (e.g., Agilent SureSelect Methyl-Seq) | Hybridization-based target capture panels designed for bisulfite-converted DNA, allowing deep sequencing of specific CpG-rich regions across many serial samples. |
| Unique Molecular Identifier (UMI) Adapters | Tag individual DNA molecules before PCR and sequencing to correct for amplification errors and generate accurate quantitative methylation measurements for tracking. |
| Methylation Reference Standards (e.g., Horizon Dx) | Commercially available DNA mixes with known methylation levels at specific loci, essential for assay validation and monitoring batch-to-batch technical variation across a longitudinal study. |
The comparative analysis of tissue and liquid biopsy methylation profiling is central to advancing precision oncology. This guide provides an objective comparison of performance characteristics, supported by experimental data, to inform strategic selection.
Performance Comparison: Tissue vs. Liquid Biopsy Methylation Analysis
The following table summarizes key performance metrics based on recent validation studies.
| Performance Metric | Tissue Biopsy | Liquid Biopsy (ctDNA) | Combined Approach |
|---|---|---|---|
| Tumor Fraction | High (≈20-80% tumor nuclei) | Low (≈0.1-10% ctDNA) | Captures both high and low fractions |
| Spatial Heterogeneity | Limited to sampled site; high resolution | Integrates tumor DNA from all sites; low resolution | Comprehensive; resolves spatial vs. systemic |
| Temporal Resolution | Single time point (invasive) | High (enables serial monitoring) | Dynamic, enables longitudinal tracking |
| Sensitivity (Detection) | N/A (direct observation) | ~90-95% for stage III-IV; lower for early stage | Maximized (>95% in advanced cancers) |
| Specificity | Very High (>99%) | High (>99% with stringent thresholds) | Very High |
| Genome Coverage | Whole methylome possible (WGBS, RRBS) | Targeted panels (50-500 genes); whole-methylome emerging | Integrative depth and breadth |
| Primary Clinical Utility | Diagnosis, histology, initial biomarker discovery | Minimal residual disease (MRD), therapy monitoring, resistance | Holistic patient management from diagnosis to relapse |
Experimental Protocols for Key Comparative Studies
Protocol 1: Paired Tissue-Plasma Analysis for Tumor-of-Origin Mapping
- Sample Collection: Collect matched FFPE tumor tissue and peripheral blood (2x10 mL Streck tubes) from the same patient.
- Tissue Processing: Macro-dissect tumor-rich areas. Extract DNA using a silica-membrane based kit (e.g., QIAamp DNA FFPE Tissue Kit). Assess quality via DIN (DNA Integrity Number).
- Plasma Processing: Double-centrifuge blood to obtain cell-free plasma. Extract cell-free DNA (cfDNA) using a high-recovery method (e.g., QIAamp Circulating Nucleic Acid Kit). Quantify using a fluorometric assay specific for short fragments.
- Bisulfite Conversion: Treat 50-500 ng of DNA from both sources with sodium bisulfite (e.g., EZ DNA Methylation-Lightning Kit) to convert unmethylated cytosines to uracil.
- Library Preparation & Sequencing: For tissue, use whole-genome bisulfite sequencing (WGBS) or targeted panels (e.g., Illumina EPIC array). For plasma, use a targeted, multiplex PCR-based or hybrid-capture panel (e.g., Guardian G360, Roche AVENIO) optimized for bisulfite-converted cfDNA. Sequence on an Illumina platform.
- Data Analysis: Align reads to a bisulfite-converted reference genome. Call methylation status at CpG sites. Use a reference database of tissue-specific methylation signatures to predict the tissue of origin for plasma-derived signals.
Protocol 2: Longitudinal Monitoring of Methylation-Based Minimal Residual Disease (MRD)
- Baseline Sample: Obtain primary tumor tissue (FFPE or fresh frozen). Perform whole-methylome profiling to identify patient-specific hypermethylated markers.
- Assay Design: Design a patient-specific ddPCR assay or a multiplex NGS panel targeting 5-10 top-ranked, highly methylated markers.
- Post-Treatment Monitoring: Collect serial plasma draws (e.g., every 3-6 months post-surgery/adjuvant therapy).
- Liquid Biopsy Processing: Isolate cfDNA and perform bisulfite conversion as in Protocol 1.
- Targeted Detection: Analyze converted cfDNA using the patient-specific assay. For ddPCR, report copies/mL of plasma for each methylated target.
- Analysis: A positive MRD signal is defined as detection of ≥2 markers above a pre-defined threshold (e.g., ≥0.01% variant allele fraction). Correlate with clinical recurrence data.
Visualizations
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Function in Methylation Analysis |
|---|---|
| Streck Cell-Free DNA BCT Tubes | Preserves blood sample by stabilizing nucleated cells, preventing genomic DNA contamination of plasma cfDNA. |
| QIAamp DNA FFPE Tissue Kit | Silica-membrane based extraction of DNA from formalin-fixed, paraffin-embedded tissue, critical for archival samples. |
| QIAamp Circulating Nucleic Acid Kit | Optimized for maximal yield of short-fragment cfDNA from large-volume plasma samples. |
| EZ DNA Methylation-Lightning Kit | Rapid, efficient sodium bisulfite conversion of DNA with minimal degradation. |
| KAPA HyperPrep Kit (with Methylation Adaptors) | Library preparation for NGS from bisulfite-converted DNA, maintaining complexity. |
| Illumina Infinium MethylationEPIC BeadChip | Array-based platform for profiling >850,000 CpG sites in tissue or high-input DNA samples. |
| Guardant G360 or Roche AVENIO ctDNA Kits | Commercially available targeted NGS panels optimized for bisulfite-converted ctDNA analysis. |
| Bio-Rad ddPCR Supermix for Probes (No dUTP) | Essential for absolute quantification of specific methylated alleles in cfDNA without bias from uracil. |
Conclusion
Tissue and liquid biopsy methylation analyses are not competing but synergistic technologies in the modern research and clinical toolkit. Tissue biopsies remain indispensable for discovering methylation signatures, understanding tumor microenvironment context, and validating novel biomarkers. Liquid biopsies, powered by advances in sensitivity and bioinformatics, excel in non-invasive longitudinal monitoring, early detection, and capturing systemic heterogeneity. The future lies in integrated epigenomic profiles, where tissue-derived biomarkers are validated and deployed in liquid assays. For drug developers, this means more robust pharmacodynamic markers and patient stratification tools. For researchers, it necessitates rigorous cross-validation of findings across matrices. Embracing both approaches will accelerate the translation of methylation science into impactful precision oncology solutions, from early interception to the management of advanced disease.